QUOKKA
Quadrilateral, Umbra-producing, Orthogonal, Kangaroo-conserving Kode for Astrophysics!
Quokka is a two-moment radiation hydrodynamics code that uses the piecewise-parabolic method, with AMR and subcycling in time. Runs on CPUs (MPI+vectorized) or NVIDIA GPUs (MPI+CUDA) with a single-source codebase. Written in C++20. (100% Fortran-free.)
Note
The Quokka methods paper is now available on arXiv.
Warning: Beta physics modules
The following modules should currently be treated as beta because they have not yet been exercised in a published science application with Quokka: MHD, radiation, dust, particles, chemistry, and self-gravity. See Known Issues and Errata for the current status and caveats.
We use the AMReX library (Zhang et al., 2019) to provide patch-based adaptive mesh functionality. We take advantage of the C++ loop abstractions in AMReX in order to run with high performance on either CPUs or GPUs.
Example simulation set-ups are included in the GitHub repository for many astrophysical problems of interest related to star formation and the interstellar medium.
Developer guides
Contact
All communication takes place on the Quokka GitHub repository. You can start a Discussion for technical support, or open an Issue for any bug reports.
About
Quokka riding a rocket
Quokka is a high-resolution shock capturing AMR radiation hydrodynamics code using the AMReX library (Zhang et al., 2019) to provide patch-based adaptive mesh functionality. We take advantage of the C++ loop abstractions in AMReX in order to run with high performance on either CPUs, NVIDIA GPUs, or AMD GPUs.
Physics module status
Per the documentation policy in the maintainer guide, physics features that have not yet been exercised in a published science application are marked as beta.
| Module | Status | Notes |
|---|---|---|
| Radiation | beta | Two-moment radiation transport and matter-radiation coupling |
| MHD | beta | Ideal MHD with constrained transport |
| Dust | beta | Dedicated dust dynamics and drag source terms |
| Particles | beta | Particle-mesh gravity, sink particles, star formation, and feedback |
| Chemistry | beta | Primordial chemistry source terms |
| Self-gravity | beta | Poisson solve for gas and particle mass |
Hydrodynamics and optically-thin cooling are not currently marked as beta.
See Known Issues and Errata for the current caveats and unsupported combinations.
Development methodology
The code is written in modern C++20, using MPI for distributed-memory parallelism, with the AMReX GPU abstraction compiling as either native CUDA code or native HIP code when GPU support is enabled.
We use a modern C++ development methodology, using CMake, CTest, and Doxygen. We use clang-format for automated code formatting, and clang-tidy and SonarCloud for static analysis, in order to audit code adherence to the ISO C++ Core Guidelines and the MISRA C/C++ guidelines. We additionally ensure the code is free of memory corruption bugs using Clang’s AddressSanitizer.
There is an automated suite of test problems that can be run using CTest. Each test problem has a validated solution against which it is compared (usually in L1 norm) in order to pass.
Code development is managed using pull requests (PRs) on GitHub. In an effort to ensure long-term code maintainability, all code must be written in C++20 following the Coding Guidelines, it must compile using Clang without warnings, all tests must pass, and the static analyzers must show zero new bugs before a pull request is merged with the main branch.
User assistance and bug reports are managed via Discussions and Issues in the GitHub repository.
Numerical methods
Hydrodynamics
The hydrodynamics solver is an unsplit method, using the piecewise parabolic method(Colella & Woodward, 1984) for reconstruction in the primitive variables, the HLLC Riemann solver (Toro, 2013) for flux computations, and a method-of-lines formulation for the time integration.
We use the method of (Miller & Colella, 2002) to reduce the order of reconstruction in zones where shocks are detected in order to suppress spurious oscillations in strong shocks.
Radiation
The radiation hydrodynamics formulation is based on the mixed-frame moment equations (e.g., (Mihalas & Mihalas, 1984)). The radiation subsystem is coupled to the hydrodynamic subsystem via operator splitting, with the hydrodynamic update computed first, followed by the radiation update, with the latter update including the source terms corresponding to the radiation four-force applied to both the radiation and hydrodynamic variables. A method-of-lines formulation is also used for the time integration, with the time integration done by the same integrator chosen for the hydrodynamic subsystem.
The hyperbolic radiation subsystem is solved using an unsplit method, using PPM for reconstruction of the moment variables, with fluxes computed via the HLL Riemann solver, with the wavespeeds computed using the ‘frozen Eddington factor’ approximation (Balsara, 1999), which is more robust than using the eigenvalues of the M1 system (Skinner & Ostriker, 2013) itself.
We reconstruct the energy density and the reduced flux \(f = F/cE\), in order to maintain the flux-limiting condition \(F \le cE\) in discontinuous and near-discontinuous radiation flows.
To ensure the correct behavior of the advection terms in the asymptotic diffusion limit (Lowrie & Morel, 2001), we modify the Riemann solver according to (Skinner et al., 2019). We use the Lorentz-factor local closure of (Levermore, 1984) to compute the variable Eddington tensor.
The source terms corresponding to matter-radiation energy exchange are solved implicitly with the method of (Howell & Greenough, 2003) following the hyperbolic subsystem update. The matter-radiation momentum update is likewise computed implicitly in order to maintain the correct behavior in the asymptotic diffusion limit (Skinner et al., 2019).
Equations
Fluids, radiation, magnetic fields, and dust
Assuming the speed of light is not reduced (\(\hat{c} = c\)), Quokka solves the following conservation laws for the cell-centered variables, as written below in the Heaviside–Lorentz system of units:
where the total fluid energy is
and the face-centered magnetic field is evolved according to the ideal MHD induction equation:
Quokka also solves the non-conservative auxiliary internal energy equation:
and the gravitational Poisson equation:
where
- \(\rho\) is the gas density,
- \(\vec{v}\) is the gas velocity,
- \(E\) is the total fluid energy density, including magnetic energy when MHD is enabled,
- \(\rho e_{\text{aux}}\) is the auxiliary gas internal energy density,
- \(X_n\) is the fractional concentration of species \(n\),
- \(\dot{X}_n\) is the chemical reaction term for species \(n\),
- \(\mathcal{H}\) is the optically-thin volumetric heating term (radiative and chemical),
- \(\mathcal{C}\) is the optically-thin volumetric cooling term (radiative and chemical),
- \(p(\rho, e)\) is the gas pressure derived from a general convex equation of state,
- \(\vec{B}\) is the magnetic field,
- \(\mathsf{I}\) is the identity tensor,
- \(E_g\) is the radiation energy density for group \(g\),
- \(F_g\) is the radiation flux for group \(g\),
- \(\mathsf{P}_g\) is the radiation pressure tensor for group \(g\),
- \(G_g\) is the radiation four-force \([G^0_g, \vec{G}_g]\) due to group \(g\),
- \(\Delta S_{\text{rad}}\) is the change in gas internal energy due to radiation over a timestep,
- \(\phi\) is the Newtonian gravitational potential,
- \(\vec{g}\) is the gravitational acceleration,
- \(\rho_i\) is the mass density due to particle \(i\),
- \(\rho_{\mathrm{d},k}\) is the dust mass density for dust species \(k\) (\(k \in [1, N_{\mathrm{dust}}]\)),
- \(\vec{v}_{\mathrm{d},k}\) is the dust velocity for dust species \(k\),
- \(T_{\mathrm{s},k}\) is the aerodynamic stopping time for dust species \(k\),
- \(\omega\) is the fraction of frictional heating deposited into the gas.
Note that since work done by radiation on the gas is included in the \(c \sum_g G^0_g\) term, \(S_{\text{rad}}\) is not the same as \(c \sum_g G^0_g\).
Collisionless particles
Quokka solves the following equation of motion for collisionless particles:
where \(\vec{x}_i\) is the position vector of particle \(i\).
Citation
If you use Quokka or numerical methods originally implemented for Quokka in your research, please cite (Wibking & Krumholz, 2022).
@ARTICLE{2022MNRAS.512.1430W,
author = {{Wibking}, Benjamin D. and {Krumholz}, Mark R.},
title = "{QUOKKA: a code for two-moment AMR radiation hydrodynamics on GPUs}",
journal = {\mnras},
keywords = {hydrodynamics, methods: numerical, Astrophysics - Instrumentation and Methods for Astrophysics},
year = 2022,
month = may,
volume = {512},
number = {1},
pages = {1430-1449},
doi = {10.1093/mnras/stac439},
archivePrefix = {arXiv},
eprint = {2110.01792},
primaryClass = {astro-ph.IM},
adsurl = {https://ui.adsabs.harvard.edu/abs/2022MNRAS.512.1430W},
adsnote = {Provided by the SAO/NASA Astrophysics Data System}
}
What to cite
Please also cite the module- or method-specific papers that apply to your work.
- Core Quokka hydro / AMR code: (Wibking & Krumholz, 2022)
- Radiation-hydrodynamics time integration: (He et al., 2024)
- Multigroup radiation-hydrodynamics: (He et al., 2024)
- Particle-mesh interactions: (He et al., 2026)
The following modules are currently marked as beta for science-use maturity in Quokka documentation: MHD, radiation, dust, particles, chemistry, and self-gravity. For beta capabilities, cite the relevant methods papers above when they exist, and report the exact Quokka version or commit hash used in your study.
Published scientific applications with Quokka
- Galactic outflows: (Vijayan et al., 2024), (Huang et al., 2025), (Vijayan et al., 2025)
Known Issues and Errata
This page collects current limitations, beta-status physics features, and any corrections to the published documentation or methods papers.
Beta physics modules
The following Quokka physics modules should currently be treated as beta because they have not yet been exercised in a published science application with Quokka:
| Module | Status | Notes | Documentation |
|---|---|---|---|
| Radiation | beta | Two-moment radiation transport and matter-radiation coupling | Equations, Radiation Integrator |
| Magnetohydrodynamics (MHD) | beta | Ideal MHD with constrained transport | MHD module |
| Dust | beta | Dedicated dust dynamics and dust-gas drag source terms | Dust module |
| Particles | beta | Particle-mesh gravity, sink particles, star formation, and feedback | Particles |
| Chemistry | beta | Primordial chemistry source terms | Equations, Runtime parameters |
| Self-gravity | beta | Poisson solve for gas and particle mass | Equations |
Hydrodynamics and optically-thin cooling are not currently marked as beta.
Current limitations
- Reflecting magnetic-field boundary conditions are not yet physically complete; Quokka currently applies
reflect_evento all magnetic-field components. MHD + radiationis not yet tested and is currently explicitly disabled in the code.MHD + dustis not exercised by an in-tree problem and should be treated as untested.- Dust currently neither contributes to, nor feels forces from, the gravitational potential.
Errata
No published errata are currently recorded on this page.
If you find a documentation error or a discrepancy between the documented equations and the implementation, please open an issue on the Quokka GitHub repository.
Glossary
Quick reference for recurring terminology across the Quokka documentation and reviews.
- PR (Pull Request): a proposed code change waiting for review.
- Revert: a follow-up change that undoes a previous merge to restore a known-good state.
- ADR (Architecture Decision Record): a short document capturing an important technical decision, options considered, and the rationale.
- CI (Continuous Integration): automated builds and tests that run on every change.
- GPU backend (CUDA/HIP): the selected GPU programming platform chosen at compile time.
- MPI: a standard for running a program across multiple processes (often across nodes).
- Unit test: a small test for a single function or module.
- Physics test (domain test): a small end-to-end run that checks scientifically meaningful results.
- Regression test: a test that ensures behavior hasn’t unintentionally changed compared to a known baseline.
- Benchmark: a timing or resource-usage measurement to track performance.
- Debug build / Release build: debug enables extra checks and slower safety features; release is optimized for speed.
- Stream (GPU): an ordered queue of GPU work; operations in a stream execute in sequence.
- Seed (random): a number used to initialize a random number generator so results can be repeated.
- Tolerance (absolute/relative): acceptable numerical error bounds (e.g., max difference allowed).
- Baseline: a trusted reference result used for comparison.
Magnetohydrodynamics Module
Warning: Beta feature
The MHD module has not yet been exercised in a published science application with Quokka and should currently be treated as beta.
The magnetohydrodynamics (MHD) module augments Quokka’s Euler solver with magnetic fields while keeping the discrete divergence of \(\vec{B}\) exactly zero. It is built on the same method-of-lines framework described in Equations, but introduces face- and edge-centred fields to support constrained transport.
Finite-volume discretization
- Cell-centred conserved variables (\(\rho\), \(\rho\vec{v}\), \(E_{\text{gas}}\) and
optional scalars) live in the same
MultiFabas the hydrodynamics module. Magnetic fluxes \(B_1\), \(B_2\), \(B_3\) are stored on face-centredMultiFabs so that each component is collocated with its normal face area. - Edge-centred electromotive forces (EMFs) are defined on a third staggered grid and assembled on demand. This staggered arrangement lets the induction update follow Stokes’ theorem exactly, ensuring \(\nabla \cdot \vec{B}=0\) to machine precision on uniform grids.
- Source terms (cooling, gravity, etc.) act on the cell-centred state. Since only ideal MHD is supported, there are no source terms in the magnetic induction equation.
Reconstruction and Riemann solves
Each sweep through computeHydroFluxes performs the following steps for every
coordinate direction:
- Convert conserved variables (plus face-centred magnetic fluxes) to primitive
variables with
HydroSystem::ConservedToPrimitive. - Build shock-flattening coefficients that detect strong compression and locally limit the reconstruction stencil to maintain monotonicity.
- Reconstruct left/right interface states using the requested order: donor-cell (1), piecewise linear with a minmod limiter (2), PPM (3) or extrema-preserving PPM (5, the default). The transverse magnetic components are reconstructed alongside the fluid primitives so that the Riemann solver sees a consistent state.
- Solve the one-dimensional Riemann problem with the HLLD solver. The normal magnetic field is taken directly from the face-centred data so that the solution respects the jump condition \(B_n^- = B_n^+\). The solver returns both fluxes for the conserved state and a face-centred velocity that is reused by particle advection and, optionally, by the EMF calculation.
If the high-order solve produces negative densities, the code marks the affected cells and later recomputes their update with a first-order upwind flux (see Fallbacks below). Negative pressures are corrected separately by the dual-energy synchronisation step.
Edge-centred electromotive forces
MHDSystem::ComputeEMF constructs either cell-centered or edge-centred EMFs \(\mathcal{E} = -\vec{v}\times\vec{B}\) by
combining face-centred magnetic fields with one of the three schemes selected by
emf_computinging_scheme:
FelkerStone2017– reconstructed velocities from cell-centered states to edges.Balsara2025– average of face-centered magnetic fields with cell-centered velocities, with subsequent EMF calculation at cell-center then reconstructed EMF to edgesQuokka2026- face-centered velocity returned by the Riemann solver reconstructed to edges
The stencil used in this reconstruction is controlled by emf_reconstruction_order
with the same order options as the flux reconstruction. After reconstruction,
Quokka averages the four quadrant-centred EMFs surrounding each edge using one of
two formulas selected by emf_averaging_scheme:
LondrilloDelZanna2004– the Londrillo & Del Zanna (2004) upwind constrained-transport formula, which weights the quadrants using characteristic MHD signal speeds.Balsara2025- EMF averaging with a dissapative term from wavespeeds and magnetic field jumps
During the LondrilloDelZanna2004 average, the code leverages the fast magnetosonic speeds computed at interfaces during the Riemann solve so that the EMF averaging is properly upwinded without requiring a full state reconstruction and eigenvalue computation.
Constrained-transport update
Once EMFs are available, MHDSystem::SolveInductionEqn updates the face-centred
magnetic fluxes using the area-integrated form of Faraday’s law. For a face with
normal direction \(w_0\) and transverse directions \(w_1\), \(w_2\) the update reads
Because the EMFs on opposite edges are equal and opposite, the discrete divergence, computed as a finite difference of the face-centred fluxes, remains unchanged by the update.
Time integration
The MHD module uses the same second-order strong stability preserving Runge–Kutta (SSPRK2) integrator that advances the hydrodynamics:
- Predictor stage: Evaluate numerical fluxes (and EMFs) from the state at
\(t^n\) and form a provisional update using
HydroSystem::PredictStep. - Corrector stage: Recompute fluxes/EMFs from the predictor state, average
them with the first-stage fluxes, and apply
HydroSystem::PredictStepa second time to obtain \(U^{n+1}\).
Strang-split source terms (chemistry, cooling, gravity) advance by half steps before and after the hydrodynamic RK stages. Radiation transport advances in a separate, fully operator-split solve after the hydrodynamic update.
Fallbacks and positivity control
- Cells flagged during reconstruction because the high-order state produced a negative density are re-updated with a first-order local Lax–Friedrichs flux and EMFs reconstructed with donor-cell stencils. This “first-order flux correction” preserves robustness while keeping the divergence constraint. (The dual-energy sync step is responsible for repairing negative pressures.)
- Density and temperature floors are enforced after every RK stage.
- When the dual-energy formalism is enabled, the auxiliary internal energy is synchronised with the magnetized total energy to avoid large truncation errors in the internal energy in high Mach number regions.
Runtime controls
The following input parameters tune the MHD discretization and are documented in more detail in Runtime parameters:
reconstruction_order– spatial order for hydro flux reconstruction (default 3 = PPM).emf_reconstruction_order– spatial order for the EMF reconstruction (default 5 = extrema-preserving PPM).emf_compute_scheme– chooseFelkerStone2017,Balsara2025, orQuokka2026for computing the emf either at cell-center or at the edge (defaultBalsara2025).emf_averaging_scheme– chooseLondrilloDelZanna2004orBalsara2025for edge averaging (defaultBalsara2025).artificial_viscosity_k– optional scalar viscosity coefficient that adds a diffusive flux to the momentum equations and can damp post-shock oscillations.quokka.bc– (required) chooseperiodicorreflectingfor the boundary conditions. For reflecting boundaries, we useamrex::BCType::reflect_evenfor all magnetic field components. Support for properly reflecting magnetic field boundaries will be added in the future.
Particles, star formation and feedback
Warning: Beta feature
The particles, star-formation, and feedback module has not yet been exercised in a published science application with Quokka and should currently be treated as beta.
All particle features can only be activated when compiled with -DAMReX_SPACEDIM=3.
Sink Particle Type
Sink particles carry the following real-valued attributes:
- Mass: Current particle mass (\(M\))
- Velocity: Three-dimensional velocity vector \((v_x, v_y, v_z)\)
Unlike StochasticStellarPop particles, sink particles have no evolution stage or birth/death metadata. They represent unresolved gravitationally-collapsed objects that accrete mass from the surrounding gas.
Sink Particle Formation
Overview
The sink formation module creates sink particles from gas cells that violate the Jeans criterion (Truelove et al. 1997). The algorithm is operator-split from the hydrodynamics and runs once per hydro timestep at the finest AMR level.
Formation criteria
A sink particle is created in a cell if all of the following conditions are met:
- Jeans instability: The cell density exceeds the local Jeans density,
where \(J = 0.25\) is the Jeans number and \(c_{\text{eff}} = c_s \sqrt{1 + 0.74 / \beta}\) is the effective sound speed accounting for magnetic pressure support (\(\beta = P_{\text{thermal}} / P_{\text{magnetic}}\)). For non-MHD simulations, \(\beta \to \infty\) and \(c_{\text{eff}} = c_s\).
-
Not in an existing accretion zone: The cell is not already being accreted by an existing sink particle (i.e., the accretion rate in the cell is zero).
-
Local density maximum: The cell has the highest density among all cells within a sphere of radius \(3 \Delta x\). This prevents multiple sink particles from forming in close proximity.
Mass and momentum assignment
When a sink particle forms, it receives the excess mass above the Jeans density:
The particle inherits the gas velocity of the parent cell. The cell density is then set to \(\rho_J\), and all conserved quantities (momentum, energy, internal energy) are scaled by the factor \(\rho_J / \rho\).
Practical considerations
- The local maximum check uses a non-strict inequality (\(\rho_{\text{cell}} \ge \rho_{\text{neighbor}}\)), so ties do not prevent formation. In practice, exact density ties only occur in artificial initial conditions.
- Sink formation is applied after sink accretion in each timestep, so newly formed particles do not overlap with existing accretion zones.
Sink Particle Accretion
Overview
Sink particle accretion follows the Bondi-Hoyle-Lyttleton prescription of Krumholz et al. (2004). Gas is removed from cells surrounding each sink particle and added to the particle’s mass, with the accretion rate set by the local gas properties and the particle mass.
Accretion rate
The Bondi-Hoyle radius is
where \(M\) is the particle mass, \(v_\infty\) is the gas velocity relative to the particle (mass-weighted mean over the accretion zone), and \(c_{f,\infty}\) is the fast magnetosonic speed. We take the upper bound of the fast speed (when the wave propagates perpendicular to the magnetic field): \(c_f^2 = c_s^2 + v_A^2 = c_s^2 (1 + 2/\beta)\). For non-MHD, this reduces to \(c_f = c_s\).
The accretion rate is
where \(\rho_\infty\) is the mean gas density in the accretion zone and \(\lambda = e^{3/2}/4\).
Accretion kernel
The accretion zone is a sphere of radius \(r_{\text{acc}} = 3 \Delta x\) centered on the particle. Within this zone, each cell receives a Gaussian weight:
where \(r\) is the distance from the particle to the cell centre. The kernel radius \(r_K\) is resolution-adaptive:
The accretion rate deposited in each cell is \(-\dot{M} , w_i / \sum_i w_i\) (negative means accreting). Optionally, a uniform kernel (\(w = 1\) for all cells in the \((7 \Delta x)^3\) stencil) can be used for testing by setting particles.sink_particle_use_uniform_kernel = 1.
Accretion limiters
Two corrections are applied to the per-cell accretion rate, in the following order:
-
Mass removal cap: No more than 25% of a cell’s mass may be removed in a single timestep (Krumholz et al. 2004). This prevents artificial sound waves from being launched by rapid density changes.
-
Jeans density floor: If the post-accretion cell density would still exceed the Jeans density \(\rho_J\), the accretion rate is increased so that the final density equals \(\rho_J\). This is safe because such cells are at the centre of highly supersonic convergence and are causally disconnected from their surroundings.
Momentum accretion
When gas is accreted, the particle gains both mass and momentum. The new particle velocity is computed from momentum conservation:
where \(\Delta m\) is the total accreted mass and \(\vec{v}{\text{gas}}\) is the mass-weighted gas velocity over the accreted cells. The gas state is then updated by scaling all conserved quantities by \((1 + \dot{M}{\text{cell}} \Delta t / (\rho V))\), which preserves the gas velocity field.
Galilean invariance
The accretion rate is computed using gas velocities in the particle frame (\(\vec{v}\infty = \vec{v}{\text{gas}} - \vec{v}_{\text{particle}}\)), ensuring Galilean invariance.
Runtime Parameters
| Parameter | Type | Default | Description |
|---|---|---|---|
particles.sink_particle_use_uniform_kernel | Boolean | 0 | Use uniform accretion kernel (for testing) |
Examples
ParticleSinkFormation Test
The ParticleSinkFormation test validates combined sink particle formation and accretion. The test checks:
- Exactly one star forms in the first timestep.
- Mass is conserved to machine precision during sink formation.
- Mass is conserved to machine precision during sink accretion.
ParticleSink Test
The ParticleSink test validates Bondi-Hoyle accretion and Galilean invariance. It runs in three phases:
- Base simulation: Runs with zero boost velocity and validates the density profile against an analytical solution.
- Boosted simulation: Runs with a boost velocity of \(10^8\) cm/s and verifies that the density profile matches the analytical solution, demonstrating Galilean invariance.
- Multi-timestep evolution: Continues the boosted simulation for additional timesteps and validates total mass conservation to machine precision.
StochasticStellarPop Particle Type
StochasticStellarPop particles carry the following real-valued attributes:
- Mass: Current particle mass (\(M_{\star}\))
- Velocity: Three-dimensional velocity vector \((v_x, v_y, v_z)\)
- Birth time: Simulation time when the particle formed
- Death time: When the particle explodes as a supernova
- Mass at birth: Initial mass before mass loss
- Luminosity: Radiation luminosity (if radiation is enabled)
Each particle also stores an integer evolution stage that tracks its lifecycle:
SNProgenitor: High-mass star (\(M > 8 M_{\odot}\)) that will explode as a supernovaSNRemnant: Compact remnant left after supernova explosionLowMassStar: Low-mass star that will not explode (not used in the current star formation implementation)LowMassComposite: Composite particle representing a population of low-mass stars
Star Formation
Overview
The star formation module adds star particles through a stochastic prescription that plugs into the Stochastic Stellar Population specialisation.
- Runs once per hydro timestep and evaluates each cell independently.
- Always spawns a
LowMassCompositeparticle when star formation is triggered. - Adds high-mass particles (\(> 8~M_{\odot}\)) probabilistically so the Chabrier (2005) initial mass function (IMF) is satisfied in expectation.
Jeans instability filter
Eligible cells are first identified through a Jeans-length check before any stochastic sampling occurs.
- Compute the Jeans density \(\rho_J = J^2 \pi c_{\text{eff}}^2 / (G \Delta x^2)\), where \(J = 0.25\) is the Jeans number and \(c_{\text{eff}} = c_s \sqrt{1 + 0.74 / \beta}\) is the effective sound speed, accounting for thermal pressure and magnetic pressure with \(\beta = P_{\text{thermal}} / P_{\text{magnetic}}\) (Mouschovias & Spitzer, 1976, ApJ, 210, 326; see also Myers et al, 2013, ApJ, 766, 97).
- Mark the cell as eligible when \(\rho > \rho_J\).
- Only eligible cells continue to the sampling steps below.
Controlling the formation rate
Two efficiency parameters tune how aggressively eligible gas is converted into star particles during each hydro step.
- \(\epsilon_{ff}\): efficiency per free-fall time, defined through \(\epsilon_{ff} = (\dot M_{\star} , t_{ff}) / (M_{cell} , \Delta t)\).
- \(\epsilon_{\star}\): fraction of the cell mass used when a particle is spawned.
- The target stellar mass for the step is \(\epsilon_{ff} M_{cell} (\Delta t / t_{ff})\).
- Bernoulli probability for spawning: \( P = \frac{\epsilon_{ff}}{\epsilon_{\star}} \frac{\Delta t}{t_{ff}} \).
- The expectation value \(\langle M_{\star} \rangle = P \epsilon_{\star} M_{cell}\) matches the target mass provided \(\Delta t < t_{ff}\); the CFL condition typically enforces that inequality.
Sampling the stellar population
Once a cell passes the filter and the Bernoulli draw succeeds, we construct the composite stellar population represented by the spawned particles.
- Every accepted draw creates one low-mass particle that represents all stars with \(M < 8 M_{\odot}\).
- High-mass stars follow the Chabrier (2005) IMF: log-normal below \(1 M_{\odot}\) and a slope of \(2.35\) above it.
- Pre-computed IMF integrals provide the mass fraction \(f_{\star,high}\) and mean mass \(\langle m \rangle_{\star,high}\) of the high-mass component.
- The expected number of massive stars is \(f_{\star,high} \epsilon_{\star} M_{cell} / \langle m \rangle_{\star,high}\); a Poisson variate with that mean sets the actual count.
- Each massive star draws its mass from the high-mass end of the IMF, while the low-mass particle retains the remaining fraction \(1 - f_{\star,high}\) of the spawned mass.
- If the Poisson draw returns zero, only the low-mass particle is inserted and it inherits the local gas velocity.
Assigning particle velocities
Sampled star particles receive velocities that combine the local bulk flow with an isotropic runaway kick.
- Each massive star draws a speed from a power-law distribution \(p(v) \propto v^{-1.8}\) truncated between 3 and 385 km s\(^{-1}\), then converts it to cgs units.
- A random direction is chosen by sampling \(\cos \theta\) uniformly in \([-1, 1]\) and \(\phi\) in \([0, 2\pi)\); the resulting kick is added to the gas velocity of the parent cell.
- The total momentum of all massive stars is accumulated, and the low-mass composite particle receives the opposite momentum so that the cell-level particle system conserves momentum.
Practical considerations
A few implementation notes help interpret corner cases and limitations of the current recipe.
- Star formation is operator-split from the hydrodynamics. When \(t_{ff}\) is unresolved (\(\Delta t \gtrsim t_{ff}\)), the true star formation rate is not captured, and this scheme provides one possible approximation; no explicit limiter is enforced beyond the CFL-controlled hydro step.
- All spawned particles are inserted at the cell centre. Other physics modules are responsible for any subsequent repositioning or feedback coupling.
Supernova Feedback
Overview
Quokka includes a particle framework for modeling stellar populations and their feedback effects on the interstellar medium (ISM). The primary feedback mechanism is supernova (SN) explosions, which inject energy and momentum into the surrounding gas. The SN module implements several physically-motivated schemes that adapt to the local resolution and density conditions.
Supernova Feedback
Physical Parameters
When a progenitor star reaches its death time, it explodes as a Type II supernova with the following canonical parameters:
| Parameter | Symbol | Value | Description |
|---|---|---|---|
| Blast energy | \(E_{\text{blast}}\) | \(10^{51}\) erg | Thermal energy injected into the ISM |
| Ejecta mass | \(m_{\text{ej}}\) | \(10 M_{\odot}\) | Mass expelled during explosion |
| Terminal momentum | \(p_{\text{snr},0}\) | \(2.8 \times 10^5 M_{\odot} , \text{km s}^{-1}\) | Asymptotic momentum of the SNR (configurable via particles.SN_p_term_Msunkmps) |
| Remnant mass | \(m_{\text{dead}}\) | \(\geq 1.4 M_{\odot}\) | Mass of the compact remnant |
| Kinetic energy | \(E_{\text{kin}}\) | \(0.5 m_{\text{ej}} v_{\text{star}}^2\) | Kinetic energy of the ejecta |
The terminal momentum is density-dependent and scales as:
where \(n_{\text{H}}\) is the ambient hydrogen number density averaged over the deposition kernel.
Deposition Kernel
SN feedback is deposited into a \((2 \times 3 + 1)^3 = 343\) cell cubic stencil centered on the particle’s location. The kernel weights are pre-computed to approximate a spherical distribution with radius \(r = 3 \Delta x\), where \(\Delta x\) is the cell width, around the host cell center. The kernel weights \(W_{ijk}\) are normalized such that \(\sum_{i,j,k} W_{ijk} = 1\).
Resolution-Adaptive Schemes
The feedback implementation uses the resolution factor \(R_M\) to determine whether the Sedov-Taylor phase is resolved:
where:
- \(M_{\text{SNR}} = M_{\text{gas}} + m_{\text{ej}}\) is the total SNR mass (gas in stencil plus ejecta)
- \(M_{\text{sf}} = 1679 M_{\odot} , n_{\text{H}}^{-0.26} \left(p_{\text{snr},0} / p_{\text{snr},0}^{\text{canonical}}\right)^2\) is the shell-formation mass, scaled so that the kinetic energy \(p_{\text{snr}}^2 / (2 M_{\text{sf}})\) is invariant when \(p_{\text{snr},0}\) is changed
When \(R_M < 1\), the Sedov-Taylor phase is resolved and the blast wave dynamics can be captured. When \(R_M \geq 1\), the resolution is insufficient and the code transitions to momentum-dominated feedback following Kim & Ostriker (2017).
Available Schemes
Quokka provides four SN feedback schemes controlled by particles.SN_scheme:
-
SN_thermal_only: Pure thermal energy injection- Deposits \(E_{\text{blast}} + E_{\text{kin}}\) into gas total energy
- Deposits ejecta mass \(m_{\text{ej}}\) and momentum \(m_{\text{ej}} \vec{v}_{\text{ej}}\) into gas momentum. No further momentum injection
- Simplest scheme, appropriate when Sedov-Taylor phase is well-resolved. Should only be used for testing.
-
Thermal and momentum schemes: The following schemes include resolution-dependent momentum injection. In these schemes, an energy of \(E_{\text{blast}}\) + \(E_{\text{kin}}\) is injected into the gas total energy, and a fraction \(f\) of the asymptotic momentum \(p_{\text{snr}}\) is injected as momentum: \(p_{\text{inject}} = f , p_{\text{snr}}\). The difference between the injected total energy and kinetic energy remains as thermal energy. Note that there is always some amount of thermal energy injected, even when \(f = 1\). The momentum fraction \(f\) is determined by the resolution factor \(R_M\) as follows:
a.
SN_thermal_or_thermal_momentum(default):- When \(R_M < 0.027\): Pure thermal (like
SN_thermal_only) with \(f = 0\) - When \(0.027 \leq R_M < 1\): Partial momentum injection with \(f = 0.529 \sqrt{R_M}\). Based on Kim & Ostriker (2017) calibration, \(f^2 = 0.28 R_M\) means that 28% of the total energy is kinetic energy in this case.
- When \(R_M \geq 1\): Full momentum injection with \(f = 1\)
b.
SN_thermal_kinetic_or_thermal_momentum:- When \(R_M < 0.027\): Pure kinetic injection with \(f = \sqrt{2 R_M}\)
- When \(0.027 \leq R_M < 1\): Partial momentum injection with \(f = 0.529 \sqrt{R_M}\)
- When \(R_M \geq 1\): Full momentum injection with \(f = 1\)
c.
SN_pure_kinetic_or_thermal_momentum:- Always injects full terminal momentum regardless of resolution with \(f = 1\)
- When \(R_M < 0.027\): Pure thermal (like
Momentum Deposition
For schemes with momentum injection, the following momentum deposition procedure guarantees Galilean invariance for a single SN event:
- Compute COM velocity of the SNR (gas + ejecta):
where \(\vec{P}_{\text{gas}}\) is the total momentum of gas in the stencil (kernel-weighted).
- Deposit momentum such that each cell receives:
where \(\vec{p}{\text{radial}} = f , p{\text{snr}} , W_{ijk} , \hat{\mathbf{r}}_{ijk}\) is the radial momentum component.
- Energy deposition includes a cross term:
The cross term accounts for the kinetic energy change from the radial expansion, ensuring Galilean invariance. This term sums to zero over all cells in the stencil, (momentum is conserved), ensuring energy conservation.
Background smoothing. To ensure the cross term cancels when summing over all cells in the stencil, the velocity field (but not the density) must be smoothed such that . In our tests, this smoothing also dramatically reduces peak velocities in SNR. In a shearing/turbulent background, this artificially homogenizes the gas velocity and changes kinetic energy unrelated to the SN. We provide a parameter particles.SN_smooth_gas_velocity=0 to turn off this smoothing: and . In this case, the cross term sums to a non-zero value: , where is the velocity of cell relative to the COM frame.
Runtime Parameters
| Parameter | Type | Default | Description |
|---|---|---|---|
particles.SN_scheme | String | SN_thermal_or_thermal_momentum | Feedback scheme (see above) |
particles.SN_p_term_Msunkmps | Float | 2.8e5 | Terminal momentum \(p_{\text{snr},0}\) in units of \(M_\odot,\mathrm{km,s}^{-1}\). The shell-formation mass \(M_\mathrm{sf}\) is scaled as \((p/p_\mathrm{canonical})^2\) to preserve the kinetic energy \(p^2/(2M_\mathrm{sf})\). |
particles.disable_SN_feedback | Boolean | 0 | Disable SN feedback entirely |
particles.verbose | Integer | 0 | Verbosity level for particle diagnostics |
particles.stellar_velocity_limit | Float | \(10^8\) cm/s | Maximum allowed stellar velocity (aborts if exceeded) |
particles.SN_smooth_gas_velocity | Integer | 1 | Smooth gas velocity in the stencil to enforce energy conservation |
Implementation Notes
Operator Splitting
SN feedback is operator-split from the hydrodynamics and applied after each timestep:
- Hydrodynamics advances the gas state by \(\Delta t\)
- Particle positions are updated by drift
- SN candidates (particles reaching death time) are identified
- Feedback is deposited into a temporary buffer
- Buffer is added to the state using a reproducibility-aware algorithm
- Particle evolution stage is updated to
SNRemnant
Numerical Reproducibility
The deposition uses a roundoff-resistant algorithm to ensure bit-for-bit reproducibility across different processor counts. The algorithm:
- Deposits feedback into a temporary buffer with a count field
- Communicates the buffer across ranks, which introduces roundoff errors due to non-associative floating-point arithmetic
- Removes low-order bits controlled by
particles.reproducibility_roundoff_redundancyto remove this error
AMR Considerations
- Feedback is always deposited at the finest level covering the particle
- If a particle sits at a coarse-fine boundary, feedback is deposited into fine-level cells (known limitation that needs to be fixed in the future)
- The reflux operation ensures conservation across AMR boundaries
- The kernel stencil size is fixed in cell widths, so the physical size adapts to local resolution
- The use of
setForceFinestLevel(true)is recommended to ensure that all SN progenitors, and thus SN events, exist at finest level.
Limitations
- Accurate radiative cooling must be present to resolve the pressure-confined snowplow phase
- The spherical kernel is approximated on a Cartesian grid, introducing mild anisotropy
- The terminal momentum formula assumes solar metallicity and Type II SNe physics
- No explicit treatment of metal enrichment (can be added via passive scalars)
Physical Motivation
The resolution-dependent momentum injection is based on the realization that under-resolved SN explosions lose energy to artificial radiative losses within a few timesteps. The momentum-based schemes compensate by directly injecting momentum when the Sedov-Taylor phase cannot be captured.
The terminal momentum \(p_{\text{snr}}\) represents the asymptotic momentum that a supernova remnant reaches in the pressure-confined snowplow phase, after most of the kinetic energy has been radiated away. This value (Kim & Ostriker 2017) is calibrated from high-resolution simulations and depends weakly on the ambient density.
The \(R_M\) factor effectively measures whether the simulation has sufficient resolution and density to capture the Sedov-Taylor expansion. When \(R_M \ll 1\), the blast wave will expand through many cells before reaching the shell-formation radius, allowing the hydrodynamics to naturally capture the momentum buildup. When \(R_M \sim 1\) or larger, the explosion is under-resolved and the code preemptively injects the expected terminal momentum.
Examples
Basic SN Test
The SN problem provides a canonical test with one SN progenitor in a uniform medium:
particles.SN_scheme = SN_thermal_or_thermal_momentum
problem.boost_vel_x = 1.0e8 # boost velocity in x direction
This test is run three times, with ambient density of \(10, 1.0\), and \(0.1 ~\mathrm{cm}^{-3}\), corresponding to a shell-formation radius of 7, 22, and 70 pc, respectively. The spatial resolution is 5 pc. The default SN_thermal_or_thermal_momentum scheme is used. The three ambient densities implies \(R_M \approx (3 \Delta x / R_{\text{sf}})^3 = 10, 0.3\), and \(0.01\), respectively, placing the scheme in the under-resolved, partially resolved, and well-resolved regimes, respectively.
In each of the three tests, the simulation is run twice, one initially at rest and one with an initial boost velocity of \(200\) km/s, to demonstrate Galilean invariance. The temperature and velocity profiles are identical in the boosted and rest frames (to some tolerance).
Random Blast Test
The RandomBlast problem provides a testbed for multiple SN explosions with various ambient conditions and bulk flows.
Dust Module
Warning: Beta feature
The dedicated dust dynamics module has not yet been exercised in a published science application with Quokka and should currently be treated as beta.
This module primarily implements two components: the dust transport term and the dust drag source term.
Equations for Gas-Dust System
where \(\omega\) controls the level of frictional heating, with \(\omega = 0\) turning it off and \(\omega = 1\) depositing all dissipation into the gas.
Variable Storage
The dust cell-centred conserved variables (\(\rho_{\mathrm{d}}\), \(\rho_{\mathrm{d}}\vec{v_{\mathrm{d}}}\)) are added to MultiFab.
Reconstruction and Riemann Solver
Dust reconstruction is performed together with gas using the same method. The Riemann Solver used is as follows:
In one dimension along the x-direction, given the left/right states \(W_d^{L/R}\), one can provide the Riemann flux for conserved variables as follows. The density flux reads (Huang & Bai 2022):
Similar expressions hold for the momentum flux for all directions.
This is implemented in src/dust/dustRiemannSolver.hpp and called in DustSystem::ComputeDustFluxes to compute the dust advection flux.
Time Integrator
A Strang-split method (Tedeschi-Prades et al. 2025) is used to integrate the dust-gas system. The Strang-split update can be expressed as:
where \(\mathcal{D}\) is the dust-gas drag operator and \(\mathcal{H}\) is the hydrodynamics operator (including both gas and dust transport). The hydrodynamics operator \(\mathcal{H}\) is handled using the explicit RK2 scheme. The drag operator \(\mathcal{D}\) is implemented in src/dust/DustDrag.hpp and called in QuokkaSimulation::addStrangSplitSourcesWithBuiltin via DustDrag::computeDustDrag.
Optional Picard iteration for dust–gas drag
Users may optionally enable Picard iteration for the dust–gas drag operator \(\mathcal{D}\). When the stopping time depends on the gas or dust velocity, enabling iteration is required to maintain an implicit dust drag update. See Runtime parameters for details.
User-defined dust stopping time
For a given problem, users must define a problem-specific dust stopping time by implementing the DustDrag::ComputeReciprocalStoppingTime function (note that this function should return the reciprocal of the stopping time). An example can be found in the src/problems/DustDamping test.
Also, users can directly use the dust stopping time calculation helper DustDrag::ComputeReciprocalStoppingTimeKwok to compute the physical dust stopping time, following Kwok (1975) with an optional supersonic correction. The stopping time of dust \(t_{\mathrm{s}}\) is given by:
When \(\gamma=1\), this expression reduces exactly to the isothermal \(t_s\). An example of its usage can be found in the src/problems/DustDampingCorrection test.
CFL Condition for Dust
For the dust-gas coupled system with N dust species, we use the following CFL condition:
Runtime Controls
The following input parameters tune the dust module and are documented in more detail in Runtime parameters:
enable_iter_stoptime– switch of iterative dust stopping time calculation.omega– controls the level of frictional heating.print_iteration_counts- switch to turn on/off printing of dust drag iteration counts for debugging.dust.density_floor- the minimum dust density value allowed in the simulation.
Gallery
AGORA disk galaxy
We simulated a Milky Way-mass disk galaxy with the AGORA initial conditions using stellar feedback from individual massive stars:
Credit: Ben Wibking / Quokka Development Team
QED III
QED is a suite of 3D hydrodynamic simulations which follow supernova-driven outflows in varying galactic environments. The QED follows outflows from a \(1\) kpc\(^2\) patch of galaxy up to a vertical height of \(\sim\) few kpc. In the third of a series of QED papers, found here, we explore how outflow properties of the outflows depend on three galactic environments: Solar neighbourhood, inner, and outer galaxy.
Solar Neighbourhood
The initial gas density and temperature profiles are derived using Solar neighbourhood gas surface density, \(13 M_{\odot}\) pc\(^{-2}\). The SN rate is \(6\times 10^{-5}\) yr\(^{-1}\) and the gas cools with Solar (top) and sub-Solar (bottom) metallicity. Though box height (\(8\) kpc) and the resolution (\(2\) pc, uniform) are the same for both the metallicities, the nature of outflows in these two cases is quite different.
The Solar metallicity case:
The sub-Solar metallicity case:
Inner Galaxy
The initial gas surface density is \(50 M_{\odot}\) pc\(^{-2}\) and the SN rate is commensurately higher, \(3\times 10^{-4}\) yr\(^{-1}\). The domain size and resolution are identical to the Solar neighbourhood case, but note how the evolution for gas cooling at Solar (top) and sub-Solar (bottom) metallicities are closer in nature than the for the Solar neighbourhood case.
The Solar metallicity case:
The sub-Solar metallicity case:
Outer Galaxy
The intial gas surface density for these runs is \(2.5 M_{\odot}\) pc\(^{-2}\) for the Outer galaxy cases. Because the inital gas surface density is lower, the gas scale height of the warm phase is much larger. To accomodate the larger gas scale height the box size is increased to \(16\) kpc in the vertical direction. The SNe go off in a relatively high density surrounding medium arresting development of large scale outflows. The mass loading factor is small.
The Solar-metallicity case looks like this:
The sub-Solar metallicity outflows look like this:
Zoom-in
\(1\) kpc zoom-in of the Solar neighbourhood case shows the spectacular rise and fall of clouds in the Solar neighbourhood!
In the Inner galaxy, the zoom-in shows how the disc seems to be breathing in and out!
QED I
The first QED paper, link here, focuses on the metal loading properties of galactic winds launched from the Solar Neighbourhood. Initially, the galaxy is metal-free. As supernovae feedback launches outflows from the galaxy, the multiphase outflows are loaded differentially with metals. The hot phase carries most of the metals while the cooler gas, which is entrained from the disc, is metal-poor. The following movie shows midplane slices of density (top), oxygen mass density (middle), and metallicity (bottom).
Installation
Building on Linux and macOS
This instruction works on both Linux and macOS. If you come across any issues on macOS, see Installation on macOS for detailed instructions.
To run Quokka, download this repository and its submodules to your local machine:
git clone --recursive https://github.com/quokka-astro/quokka.git
Quokka uses CMake (and optionally, Ninja) as its build system. If you don’t have CMake and Ninja installed, the easiest way to install them is to run:
python3 -m pip install cmake ninja --user
Alternatively, if you have uv installed, you can use:
uv pip install cmake ninja
Now that CMake is installed, create a build/ subdirectory and compile Quokka, as shown below.
cd quokka
mkdir build; cd build
cmake .. -DCMAKE_BUILD_TYPE=Release -G Ninja
ninja -j6
Congratuations! You have now built all of the 1D test problems on CPU. You can run the automated test suite:
ninja test
You should see output that indicates all tests have passed, like this:
100% tests passed, 0 tests failed out of 20
Total Test time (real) = 111.74 sec
To run in 2D or 3D, build with the -DAMReX_SPACEDIM CMake option, for example:
cmake .. -DCMAKE_BUILD_TYPE=Release -DAMReX_SPACEDIM=3 -G Ninja
ninja -j6
to compile Quokka for 3D problems.
By default, Quokka compiles itself only for CPUs. If you want to run Quokka on GPUs, see the section “Running on GPUs” below.
Have fun!
Building with CMake + make
If you are unable to install Ninja, you can instead use CMake with the Makefile generator, which should produce identical results but is slower:
cmake .. -DCMAKE_BUILD_TYPE=Release -G "Unix Makefiles"
make -j6
make test
Could NOT find Python error
If CMake prints an error saying that Python could not be found, e.g.:
-- Could NOT find Python (missing: Python_EXECUTABLE Python_INCLUDE_DIRS Python_LIBRARIES Python_NumPy_INCLUDE_DIRS Interpreter Development NumPy Development.Module Development.Embed)
you should be able to fix this by installing NumPy (and matplotlib) by running
python3 -m pip install numpy matplotlib --user
or with uv:
uv pip install numpy matplotlib
This should enable CMake to find the NumPy header files that are needed to successfully compile.
Alternatively, you can work around this problem by disabling Python support. Python and NumPy are only used to plot the results of some test problems, so this does not otherwise affect Quokka’s functionality. Add the option
-DQUOKKA_PYTHON=OFF
to the CMake command-line options (or change the QUOKKA_PYTHON option to OFF in CMakeLists.txt).
Running on GPUs
By default, Quokka compiles itself to run only on CPUs. Quokka can run on either NVIDIA or AMD GPUs. Consult the sub-sections below for the build instructions for a given GPU vendor.
NVIDIA GPUs
If you want to run on NVIDIA GPUs, re-build Quokka as shown below. (CUDA >= 11.7 is required. Quokka is only supported on Volta V100 GPUs or newer models. Your MPI library must support CUDA-aware MPI.)
cmake .. -DCMAKE_BUILD_TYPE=Release -DAMReX_GPU_BACKEND=CUDA -DAMReX_SPACEDIM=3 -G Ninja
ninja -j6
All GPUs on a node must be visible from each MPI rank on the node for efficient GPU-aware MPI communication to take place via CUDA IPC. When using the SLURM job scheduler, this means that --gpu-bind should be set to none.
The compiled test problems are in the test problem subdirectories in build/src/. Example scripts for running Quokka on compute clusters are in the scripts/ subdirectory.
Note that 1D problems can run very slowly on GPUs due to a lack of sufficient parallelism. To run the test suite in a reasonable amount of time, you may wish to exclude the matter-energy exchange tests, e.g.:
ctest -E "MatterEnergyExchange*"
which should end with output similar to the following:
100% tests passed, 0 tests failed out of 18
Total Test time (real) = 353.77 sec
AMD GPUs
Requires ROCm 6.3.0 or newer. The directory containing the HIP and other related binaries must be added to the
PATHenvironment variable after the ROCm installation.
Build with -DAMReX_GPU_BACKEND=HIP. Your MPI library must support GPU-aware MPI for AMD GPUs. The typical AMD GPU compilers are amdclang++ or hipcc. In case your GPU-aware compiler is not being used by default during the build, use the DCMAKE_CXX_COMPILER and DCMAKE_C_COMPILER options to specify the C++ and C compilers respectively. Additionally, the AMD GPU architecture may have to be specified. This can be done using the DAMReX_GPU_ARCH option. The GPU architecture can be found using
rocminfo | grep gfx
A typical build command using the amdclang++ compiler and an AMD GPU with RDNA 2 (gfx1031) architecture will look like
cmake .. -DCMAKE_BUILD_TYPE=Release \
-DCMAKE_CXX_COMPILER=amdclang++ \
-DCMAKE_C_COMPILER=amdclang \
-DAMReX_GPU_BACKEND=HIP \
-DAMReX_GPU_ARCH=gfx1031 \
-G Ninja
Quokka has been tested on MI100, MI250X and 6700XT GPUs.
Intel GPUs (does not compile)
Due to limitations in the Intel GPU programming model, Quokka currently cannot be compiled for Intel GPUs. (See https://github.com/quokka-astro/quokka/issues/619 for the technical details.)
Building a specific test problem
By default, all available test problems will be compiled. If you only want to build a specific problem, you can list all of the available CMake targets:
cmake --build . --target help
and then build the problem of interest:
ninja -j6 test_hydro3d_blast
Building on macOS
This guide provides detailed instructions for building Quokka on macOS systems.
Prerequisites
Before installing Quokka, you need to ensure that you have a working C++ compiler, MPI library, CMake, Ninja, and HDF5 installed on your system.
Step 1: Verify C++ Compiler
First, check if you have a working C++ compiler installed by compiling a simple program:
cat > /tmp/cpp.cpp <<'EOF'
#include <iostream>
int main(){ std::cout << "C++ works\n"; }
EOF
clang++ /tmp/cpp.cpp -o /tmp/cpp && /tmp/cpp
If this command succeeds and prints “C++ works”, you’re good to go. If not, you’ll need to install Xcode Command Line Tools:
xcode-select --install
Follow the prompts to complete the installation, then verify C++ works again using the test above.
Step 2: Verify and Install MPI
Check if MPI is already installed:
mpicxx --version
If MPI is not installed, install it using Homebrew:
# Install Homebrew if you don't have it
/bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
# Install Open MPI
brew install open-mpi
After installation, verify that MPI works correctly:
mpicxx --version
mpicxx --show
cat > /tmp/mpi_cpp.cpp <<'EOF'
#include <mpi.h>
#include <iostream>
int main(int argc,char**argv){
MPI_Init(&argc,&argv);
int r; MPI_Comm_rank(MPI_COMM_WORLD,&r);
std::cout<<"Hello from C++ rank "<<r<<"\n";
MPI_Finalize();
return 0;
}
EOF
mpicxx /tmp/mpi_cpp.cpp -o /tmp/mpi_cpp && /tmp/mpi_cpp
This should compile and run successfully, printing “Hello from C++ rank 0”.
Step 3: Install CMake and Ninja
You can install CMake and Ninja using either pip or Homebrew.
Option 1: Install via pip or uv
Using pip:
python3 -m pip install cmake ninja --user
Using uv (if you have uv installed):
uv tool install cmake ninja
Note: For uv, you may also use uv pip install cmake ninja if you’re working within a virtual environment.
Option 2: Install via Homebrew
brew install cmake ninja
Verify the installation:
cmake --version
ninja --version
Step 4: Install HDF5 (Required)
HDF5 is required to build Quokka. On macOS, we recommend installing it with Homebrew:
brew install hdf5
Step 5: Install Python Dependencies (Optional but Recommended)
Some test problems use Python for plotting results. Install NumPy and matplotlib:
Using pip:
python3 -m pip install numpy matplotlib --user
Using uv:
uv pip install numpy matplotlib
Set PYTHONPATH to the site-packages directory of your Python environment. For example:
export PYTHONPATH=~/softwares/quokka/.venv/lib/python3.14/site-packages
If PYTHONPATH is unset or points to the wrong environment, post-processing imports may fail at runtime (for example ModuleNotFoundError: No module named 'numpy' or ImportError: numpy._core.multiarray failed to import), and tests such as RadDust can abort when loading matplotlib.
If you skip this step, you can disable Python support later by adding -DQUOKKA_PYTHON=OFF to the CMake configuration.
Building Quokka
Continue with the instructions in Building on Linux and macOS.
Troubleshooting
MPI Compiler Issues
If you encounter issues with the MPI compiler, you can explicitly specify it:
cmake .. -DCMAKE_CXX_COMPILER=mpicxx -DCMAKE_C_COMPILER=mpicc -DCMAKE_BUILD_TYPE=Release -G Ninja
Running on HPC clusters
Instructions for running on various HPC clusters are given below.
Gadi (NCI Australia)
The recommended build procedure on Gadi is:
source scripts/hpc_profiles/gadi_hopper.profile
cmake -B build -S . -DCMAKE_BUILD_TYPE=Release -DAMReX_GPU_BACKEND=CUDA -DAMReX_SPACEDIM=3
cmake --build build -j 8 --target HydroBlast3D
Then a single-node test job can be run with:
qsub scripts/pbs/gpuhopper-1node.pbs
You can replace HydroBlast3D with the name of the problem you want to compile.
Using VisIt
You can use VisIt in client/server mode with the following server-side patch for the launcher script.
A host file is provided here. You must change the username, project code, and server-side VisIt path.
Setonix (Pawsey)
The recommended build procedure on Setonix is:
source scripts/hpc_profiles/setonix-gpu.profile
cmake -B build -S . -DCMAKE_BUILD_TYPE=Release -DAMReX_GPU_BACKEND=HIP -DAMReX_SPACEDIM=3 -DHDF5_ROOT="$PAWSEY_HDF5_HOME"
cmake --build build -j 32 --target HydroBlast3D
Then a single-node test job can be run with:
sbatch scripts/slurm/setonix-1node.submit
You can replace HydroBlast3D with the name of the problem you want to compile.
Workaround for interconnect issues
If interconnect issues are observed, it is recommended to add the line :
export FI_CXI_RX_MATCH_MODE=software
to your job scripts.
Adaptive Mesh Refinement Guidelines
Quokka relies on the Berger-Collela AMR algorithm (Berger & Colella, 1989) provided by AMReX to create a hierarchy of nested grids. This page distills practical advice for planning and tuning AMR runs so the mesh tracks the physics you care about without wasting resolution elsewhere.
Understanding the Refinement Workflow
AMReX walks through a predictable sequence each time it builds or adjusts the mesh hierarchy. Cells are tagged inside ErrorEst, those tags are coarsened according to blocking_factor/ref_ratio, clustered by the Berger-Rigoutsos algorithm (Berger & Rigoutsos, 1991), and then promoted to new grids that honor blocking_factor and max_grid_size. The resulting hierarchy is averaged down so coarse levels remain consistent with fine ones. Keeping this loop in mind helps interpret why the mesh responds the way it does when the problem evolves or when you revisit your refinement logic.
In Quokka, AMReX still calls an ErrorEst function, but this is now a thin wrapper around refineGrid that also invokes refineGridsAroundParticles to tag cells surrounding particles. The user should therefore define refineGrid instead of ErrorEst; the wrapper will automatically call refineGrid and then add particle-based tags on top.
Setting AMR Parameters
Several inputs shape how aggressively the hierarchy responds to refinement tags:
| Parameter | Description | Typical Values | Performance Impact |
|---|---|---|---|
blocking_factor | Minimum grid dimension; grids must be integer multiples of this value | 8-32 | Large values improve GPU throughput but can force broad refinement |
grid_efficiency | Minimum fraction of tagged cells in each grid | 0.7-1.0 | Higher values reduce wasted work yet may spawn extra boxes |
max_grid_size | Maximum dimension of a grid patch | 64-128 | Smaller sizes improve load balance; larger sizes cut communication |
ref_ratio | Refinement ratio between adjacent levels | 2 or 4 | Larger ratios sharpen features faster but deepen the hierarchy |
Choose these controls as a set: a generous blocking_factor demands tighter tagging or higher grid_efficiency to avoid carpet refinement, while a smaller blocking factor allows surgical meshes but can stress GPUs. When in doubt, run short experiments that sweep one parameter at a time and compare the resulting plotfile headers to see how many grids are created per level.
Performance and Scaling Considerations
GPU runs typically favor blocking_factor ≥ 32 so each patch keeps SMs busy; values below 32 almost always break memory coalescing across a warp and leave expensive kernels underutilized even when occupancy looks fine. CPU multigrid solves prefer at least 8. If you push those limits, guard your memory footprint by trimming the number of refinement levels or enlarging max_grid_size so patches do not proliferate. Also watch how frequently you trigger regrid calls—rapid regridding magnifies the cost of tag evaluation and repartitioning, so coarse control logic or hysteresis inside refineGrid can pay dividends on very dynamic problems.
Observing and Adjusting the Hierarchy
Inspect the grid structure early in a run by opening the plotfile header or by enabling verbose regridding logs in AMReX. When the layout diverges from expectations, compare the tagged volume to the resulting grid inventory—discrepancies usually trace back to either tag dilution during coarsening or to efficiency limits that extend each patch. The spherical collapse study documented in GitHub issue #978 is a useful reminder that large blocking factors can still trigger whole-domain refinement if tags are sparse, so monitor that scenario whenever you upscale a production run.
Inspecting grids with amrex_fboxinfo
AMReX ships a lightweight reporter named amrex_fboxinfo that prints the box layout stored in a plotfile. Make sure your CMake cache was configured with -DAMReX_PLOTFILE_TOOLS=ON, then build the utility once per build tree with cmake --build build --target fboxinfo. The resulting executable lives at build/extern/amrex/Tools/Plotfile/amrex_fboxinfo. Point it at any plotfile to see how much of the domain each AMR level occupies and how many boxes AMReX created:
$ build/extern/amrex/Tools/Plotfile/amrex_fboxinfo plt00040
plotfile: plt00040
level 0: number of boxes = 1, volume = 100.00%, number of cells = 16384
maximum zones = 128 x 128
level 1: number of boxes = 4, volume = 12.50%, number of cells = 8192
maximum zones = 256 x 256
The summary lines are the quickest health check: volume reports the percentage of the level-ℓ domain covered by refined grids, so values creeping toward 100% warn that your refinement criteria are effectively forcing a uniform mesh. The maximum zones line prints the physical extents of each level’s valid domain; large jumps confirm that refinement ratios and blocking factors coarsen or refine by the factors you expect. Add --full to list every box and verify the coordinates land on multiples of blocking_factor, or --gridfile to emit a grid-description file for AMReX regression tools. When you only need the number of active refinement levels (for log parsing, for example), pass --levels to suppress the rest of the report.
Example: Berger-Rigoutsos in Practice
This figure shows an illustrative 2D blast-wave tagging pattern. The setup uses amr.n_cell = 128 128 4, amr.ref_ratio = 2 2 1, amr.blocking_factor = 32 32 4, amr.grid_eff = 0.8, and amr.n_error_buf = 1; in a 2D build the third entry in each list is ignored, so the effective blocking factor is 32. A pressure-gradient trigger in refineGrid tags a thin ring around the blast front. AMReX coarsens those tags by 16 cells before clustering, so the Berger-Rigoutsos pass finds four rectangles that tile the annulus: two long bands (lo, hi) = (64,96)→(191,127) and (64,128)→(191,159) plus two shorter bridges (96,64)→(159,95) and (96,160)→(159,191). Running amrex_fboxinfo --full plt00040 on the resulting plotfile shows the same coordinates, confirming that the level-one patches are multiples of 32 cells in each direction and fully envelop the tagged region after refinement. Comparing those boxes with the tags in the plotfile, as sketched above, is a quick way to verify that the clustering step is behaving the way you expect.
Troubleshooting Patterns
Start with the simplest levers. Reduce blocking_factor temporarily to verify the tagging logic, then restore the larger production value once you confirm the tags are reasonable. Next, nudge grid_efficiency upward until the refined region stabilizes without spawning many more boxes. If the hierarchy thrashes between steps, consider caching diagnostic arrays that record why cells were tagged; a quick visualization of those masks often pinpoints whether gradients, thresholds, or boundary effects are misbehaving.
Further Reading
For deeper dives into refinement theory and AMReX implementation details, consult the original Berger & Colella paper (Berger & Colella, 1989), the Berger-Rigoutsos clustering method (Berger & Rigoutsos, 1991), and the AMReX-Astro Castro regridding notes. The AMReX source documentation also clarifies lower-level options that Quokka exposes through its input files.
Test problems
Click here to view a table of all test problems in the Quokka codebase.
Listed here are detailed descriptions of some of the test problems. This list is still under construction.
- A table of all test problems
- Radiative shock test
- Shu-Osher shock test
- Slow-moving shock test
- Matter-radiation temperature equilibrium test
- Uniform advecting radiation in diffusive limit
- Advecting radiation pulse test
Table of all test problems
This table lists all test problems in the Quokka codebase. The acronyms used are as follows:
- SG: Single-group radiation
- MG: Multi-group radiation
- ThermalDust: Dust thermally coupled to the gas
- PE: Photoelectric heating
- CIC: Cloud-in-cell particles
| Problem | DIM | Hydro | MHD | Rad | Gravity | Particles | PassiveScalars |
|---|---|---|---|---|---|---|---|
| Advection | 1 | ❌ | ❌ | ❌ | ❌ | ❌ | ❌ |
| AdvectionSemiellipse | 1 | ❌ | ❌ | ❌ | ❌ | ❌ | ❌ |
| AlfvenWaveCircular | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| AlfvenWaveLinear | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| BinaryOrbitCIC | 3 | ✅ | ❌ | ❌ | ✅ | CIC | ❌ |
| BrioWuShockTube | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| CurrentSheet | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| DiskGalaxy | 3 | ✅ | ❌ | ❌ | ✅ | CIC, StochasticStellarPop | ❌ |
| FCQuantities | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| FastWave | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| FieldLoop | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| GravRadParticle3D | 3 | ❌ | ❌ | SG | ✅ | CIC, Rad, CICRad | ❌ |
| HydroBlast3D | 3 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroContact | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | 2 |
| HydroHighMach | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroLeblanc | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroQuirk | 2 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroSMS | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroShocktube | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroShocktubeCMA | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | 3 |
| HydroShuOsher | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroVacuum | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroWave | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| HydroWaveConvergence | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| MHDBlast | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| MHDQuirk | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| NscbcChannel | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | 1 |
| NscbcVortex | 2 | ✅ | ❌ | ❌ | ❌ | ❌ | 1 |
| ODEIntegration | 1 | ❌ | ❌ | ❌ | ❌ | ❌ | ❌ |
| OrszagTang | 3 | ✅ | ✅ | ❌ | ❌ | ❌ | ❌ |
| ParticleAccretion | 3 | ✅ | ❌ | ❌ | ✅ | Sink | ❌ |
| ParticleCreation | 3 | ✅ | ❌ | ❌ | ✅ | Test | ❌ |
| ParticleRadiation | 3 | ✅ | ❌ | MG | ❌ | StochasticStellarPop | ❌ |
| ParticleSF | 3 | ✅ | ❌ | ❌ | ❌ | StochasticStellarPop | ❌ |
| ParticleSink | 3 | ✅ | ❌ | ❌ | ✅ | Sink | ❌ |
| ParticleSinkFormation | 3 | ✅ | ❌ | ❌ | ✅ | Sink | ❌ |
| PassiveScalar | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | 1 |
| PopIII | 3 | ✅ | ❌ | ❌ | ✅ | ❌ | ❌ |
| PrimordialChem | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| RadDust | 1 | ✅ | ❌ | SG+ThermalDust | ❌ | ❌ | ❌ |
| RadDustMG | 1 | ✅ | ❌ | MG+ThermalDust | ❌ | ❌ | ❌ |
| RadForce | 1 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RadLineCooling | 1 | ✅ | ❌ | SG+ThermalDust | ❌ | ❌ | ❌ |
| RadLineCoolingMG | 1 | ✅ | ❌ | MG+ThermalDust+PE | ❌ | ❌ | ❌ |
| RadMarshak | 1 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadMarshakAsymptotic | 1 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadMarshakCGS | 1 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadMarshakDust | 1 | ❌ | ❌ | MG+ThermalDust | ❌ | ❌ | ❌ |
| RadMarshakDustPE | 1 | ❌ | ❌ | MG+ThermalDust+PE | ❌ | ❌ | ❌ |
| RadMarshakVaytet | 1 | ❌ | ❌ | MG | ❌ | ❌ | ❌ |
| RadMatterCoupling | 1 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadMatterCouplingRSLA | 1 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadStreaming | 1 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadStreamingY | 2 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadSuOlson | 1 | ❌ | ❌ | SG | ❌ | ❌ | ❌ |
| RadTube | 1 | ✅ | ❌ | MG | ❌ | ❌ | ❌ |
| RadhydroBB | 1 | ✅ | ❌ | MG | ❌ | ❌ | ❌ |
| RadhydroPulse | 1 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RadhydroPulseDyn | 1 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RadhydroPulseGrey | 1 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RadhydroPulseMGconst | 1 | ✅ | ❌ | MG | ❌ | ❌ | ❌ |
| RadhydroPulseMGint | 1 | ✅ | ❌ | MG | ❌ | ❌ | ❌ |
| RadhydroShell | 3 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RadhydroShock | 1 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RadhydroShockCGS | 1 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RadhydroShockMultigroup | 1 | ✅ | ❌ | MG | ❌ | ❌ | ❌ |
| RadhydroUniformAdvecting | 1 | ✅ | ❌ | SG | ❌ | ❌ | ❌ |
| RandomBlast | 3 | ✅ | ❌ | ❌ | ✅ | ❌ | 1 |
| RayleighTaylor3D | 3 | ✅ | ❌ | ❌ | ❌ | ❌ | 1 |
| ResampledCoolingTest | 1 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
| SN | 3 | ✅ | ❌ | ❌ | ❌ | Test | ❌ |
| ShockCloud | 3 | ✅ | ❌ | ❌ | ❌ | ❌ | 3 |
| SphericalCollapse | 3 | ✅ | ❌ | ❌ | ✅ | CIC | ❌ |
| StarCluster | 3 | ✅ | ❌ | ❌ | ✅ | ❌ | ❌ |
| Turbulence | 3 | ✅ | ❌ | ❌ | ❌ | ❌ | ❌ |
Radiative shock test
This test problem demonstrates the correct coupled solution of the hydrodynamics and radiation moment equations for a subcritical radiative shock. The steady-state solution in the nonequilibrium radiation diffusion approximation is given by a set of coupled ODEs that can be solved to arbitrary precision following the method of (Lowrie & Edwards, 2008).
Parameters
The dimensionless shock parameters (Lowrie & Edwards, 2008) are:
Following (Skinner et al., 2019), we scale to dimensional values assuming
and obtain the following pre-shock and post-shock states:
We adopt a reduced speed of light (as used in (Skinner et al., 2019))
Solution
Since the solution is given assuming radiation diffusion, we set the Eddington factor (as used in the Riemann solver for the radiation moment equations) to a constant value of \(1/3\) everywhere.
We use the RK2 integrator with a CFL number of 0.2 and a mesh of 256 equally-spaced zones. After 3 shock crossing times, we obtain a solution for the radiation temperature and matter temperature that agrees to better than 0.5% (in relative L1 norm) with the steady-state ODE solution to the radiation hydrodynamics equations:
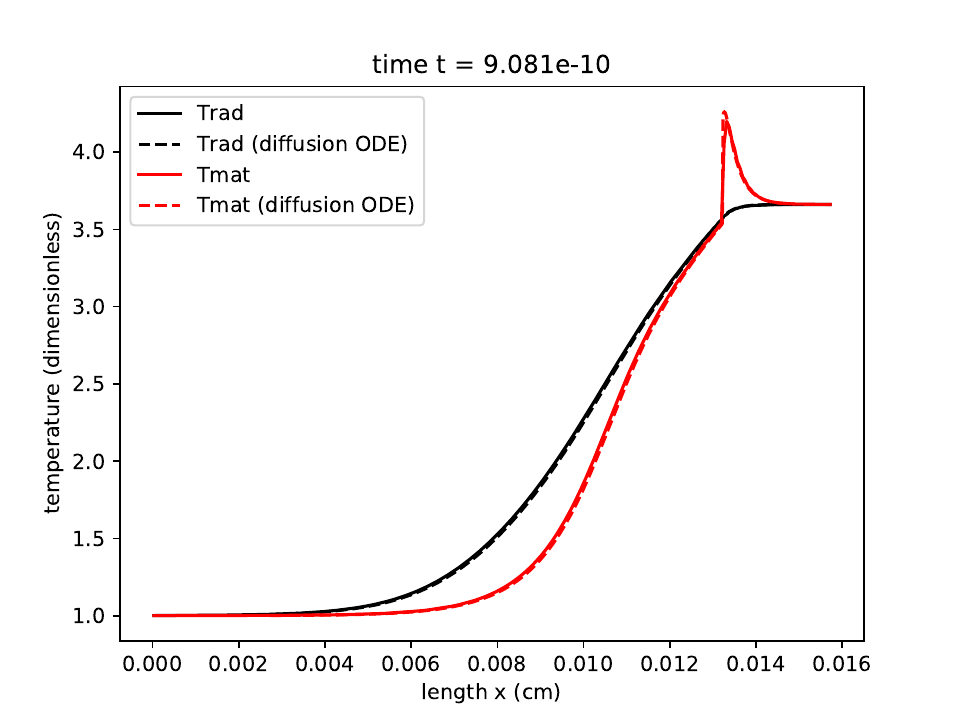 The radiation temperature is shown in the black solid and dashed lines, with the dashed line showing the semi-analytic solution. The material temperature is shown in the red lines, with the semi-analytic solution shown with the dashed line.
The radiation temperature is shown in the black solid and dashed lines, with the dashed line showing the semi-analytic solution. The material temperature is shown in the red lines, with the semi-analytic solution shown with the dashed line.
Shu-Osher shock test
This test problem demonstrates the ability of the code to resolve fine features and shocks simultaneously. The resolution of our code is comparable to the ENO-RF-3 scheme shown in Figure 14b of (Shu & Osher, 1989).
Parameters
The left- (\(x < 1.0\)) and right-side (\(x \ge 1.0\)) initial conditions are:
Solution
We use the RK2 integrator with a fixed timestep of \(10^{-4}\) and a mesh of 400 equally-spaced zones, evolving until time \(t=1.8\). There are some subtle stair-step artifacts similar to those seen in the sawtooth linear advection test, but these converge away as the spatial resolution is increased.
These artefacts can be eliminated by projecting into characteristic waves and reconstructing the interface states in the characteristic variables, as done in §4 of (Shu & Osher, 1989). The reference solution is computed using Athena++ with PPM reconstruction in the characteristic variables on a grid of 1600 zones.
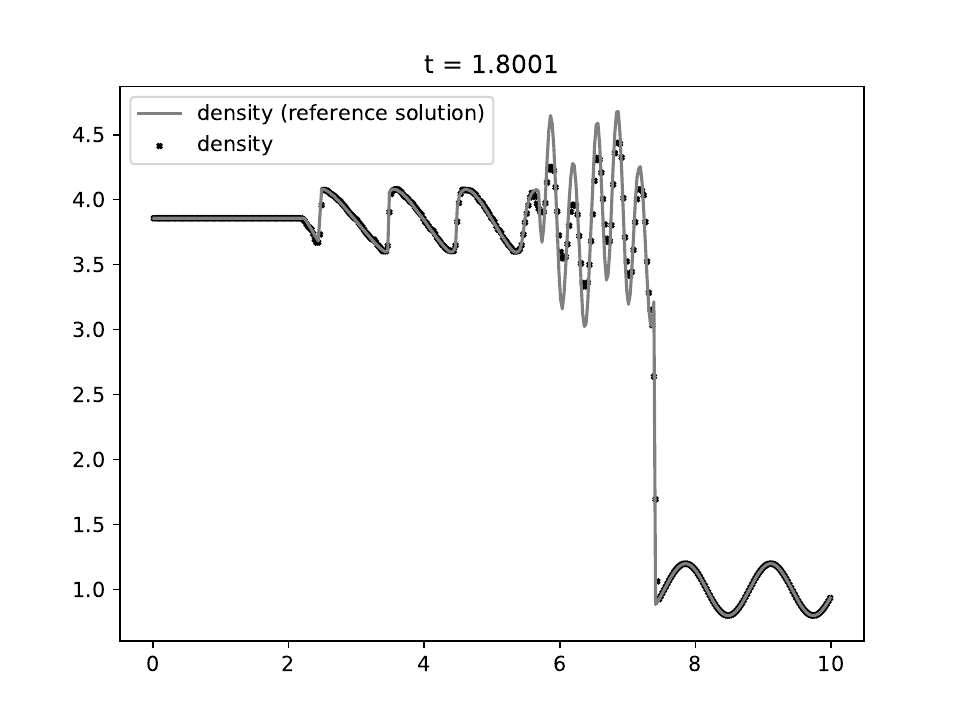 The density is shown as the solid blue line. There is no exact solution for this problem.
The density is shown as the solid blue line. There is no exact solution for this problem.
Slow-moving shock test
This test problem demonstrates the extent to which post-shock oscillations are controlled in slowly-moving shocks. This effect can be exhibited in all Godunov codes, even with first-order methods, for sufficiently slow-moving shocks across the computational grid (Jin & Liu, 1996).
The shock flattening method of (Colella & Woodward, 1984) (implemented in our code in modified form) reduces the oscillations, but does not completely suppress them. Adding artificial viscosity according to the method of (Colella & Woodward, 1984), even to the level of smoothing the contact discontinuity by 5-10 cells, does not cure the problem.
Parameters
The left- and right-side initial conditions are (Quirk, 1994):
The shock moves to the right with speed \(s = 0.1096\).
Solution
We use the RK2 integrator with a fixed timestep of \(10^{-3}\) and a mesh of 100 equally-spaced cells. The contact discontinuity is initially placed at \(x=0.5\).
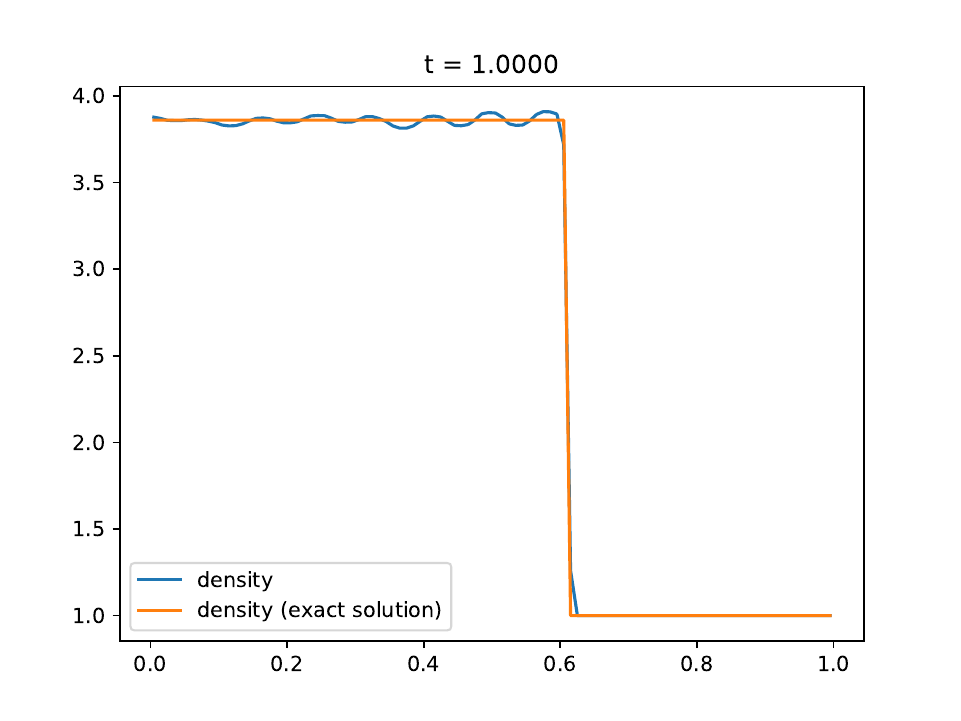 The density is shown as the solid blue line. The exact solution is the solid orange line.
The density is shown as the solid blue line. The exact solution is the solid orange line.
Matter-radiation temperature equilibrium test
This test problem demonstrates the correct coupled solution of the matter-radiation energy balance equations. We also demonstrate that the equilibrium temperature is incorrect in the reduced speed of light approximation.
Parameters
The initial energy densities are:
We assume a specific heat \(c_v = \alpha T^3\) which enables an analytic solution. We adopt a reduced speed of light with \(\hat c = 0.1 c\).
Solution
The exact time-dependent solution for the matter temperature \(T\) is:
We show the numerical results below:
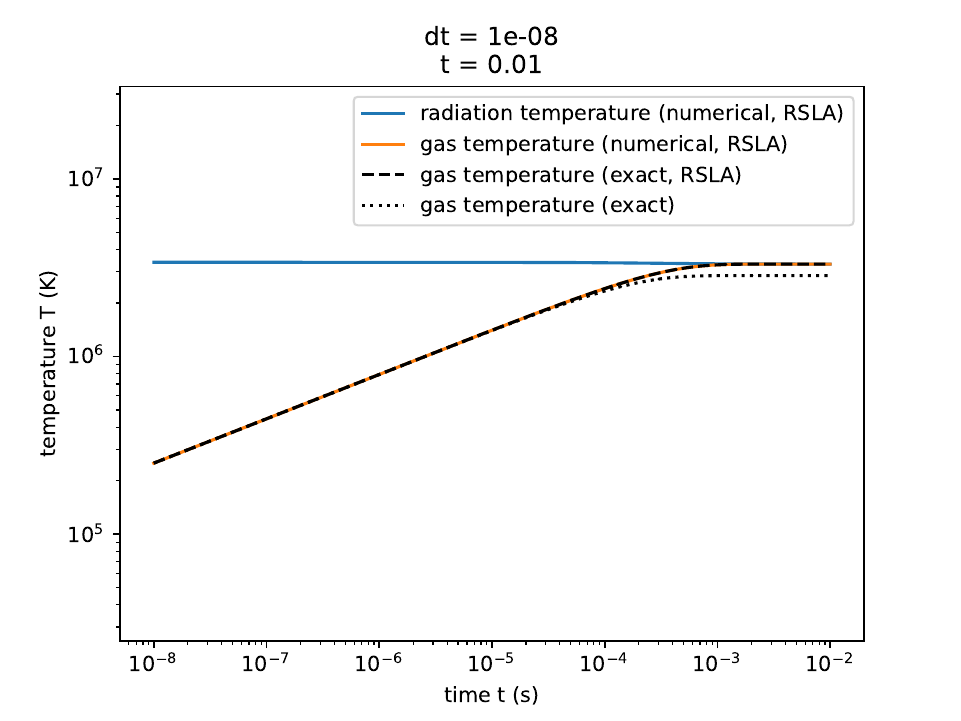 The radiation temperature and matter temperatures in the reduced speed-of-light approximation, along with the exact solution for the matter temperature.
The radiation temperature and matter temperatures in the reduced speed-of-light approximation, along with the exact solution for the matter temperature.
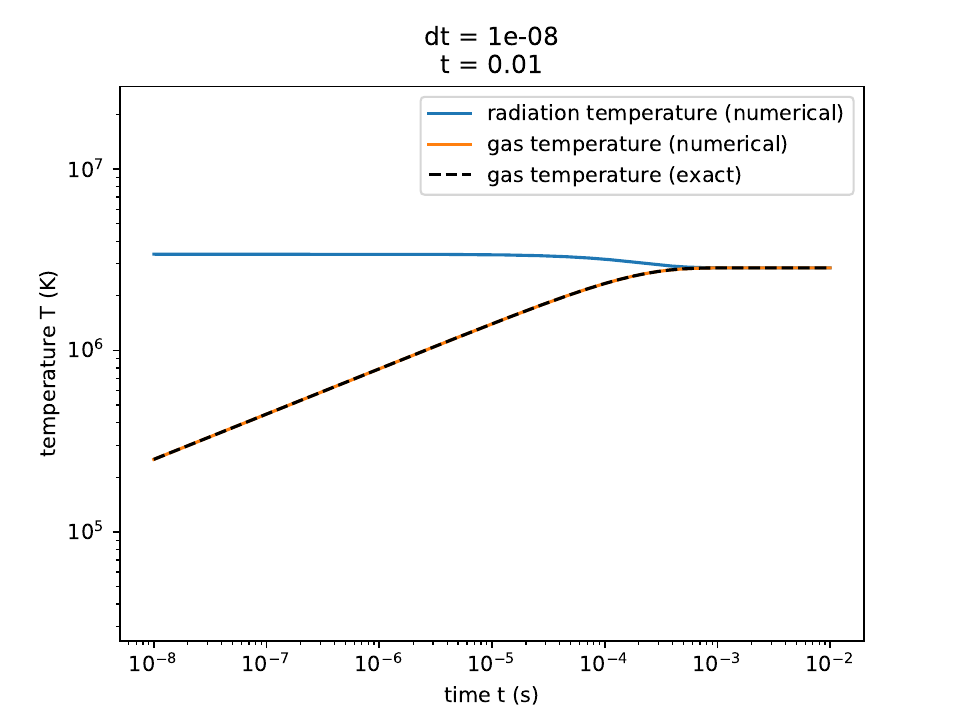 The radiation temperature and matter temperatures, along with the exact solution for the matter temperature.
The radiation temperature and matter temperatures, along with the exact solution for the matter temperature.
Uniform advecting radiation in diffusive limit
In this test, we simulation an advecting uniform gas where radiation and matter are in thermal equilibrium in the co-moving frame. Following the Lorentz tranform, the initial radiation energy and flux in the lab frame to first order in \(v/c\) are \(E_r = a_r T^4\) and \(F_r = \frac{4}{3} v E_r\).
Parameters
Results
With \(O(\beta \tau)\) terms:
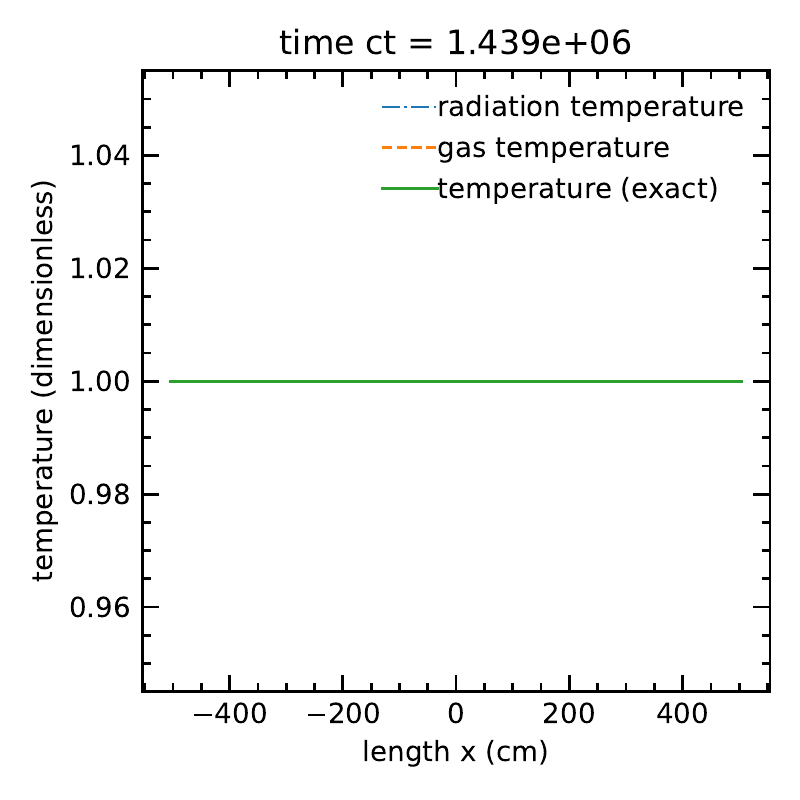 The radiation temperature and matter temperatures, along with the exact solution.
The radiation temperature and matter temperatures, along with the exact solution.
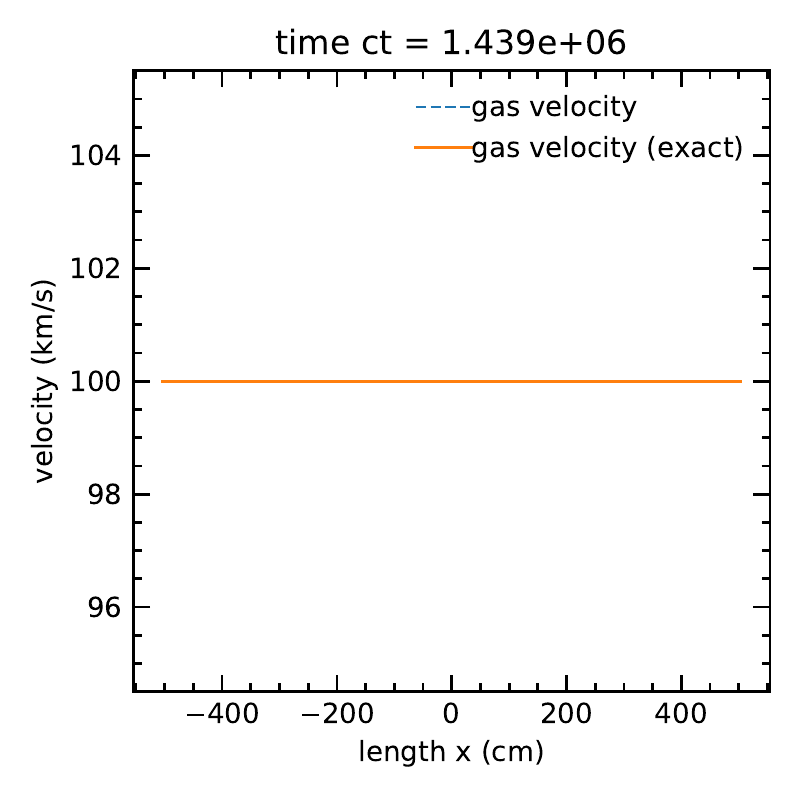 The matter velocity, along with the exact solution.
The matter velocity, along with the exact solution.
Without \(O(\beta \tau)\) terms:
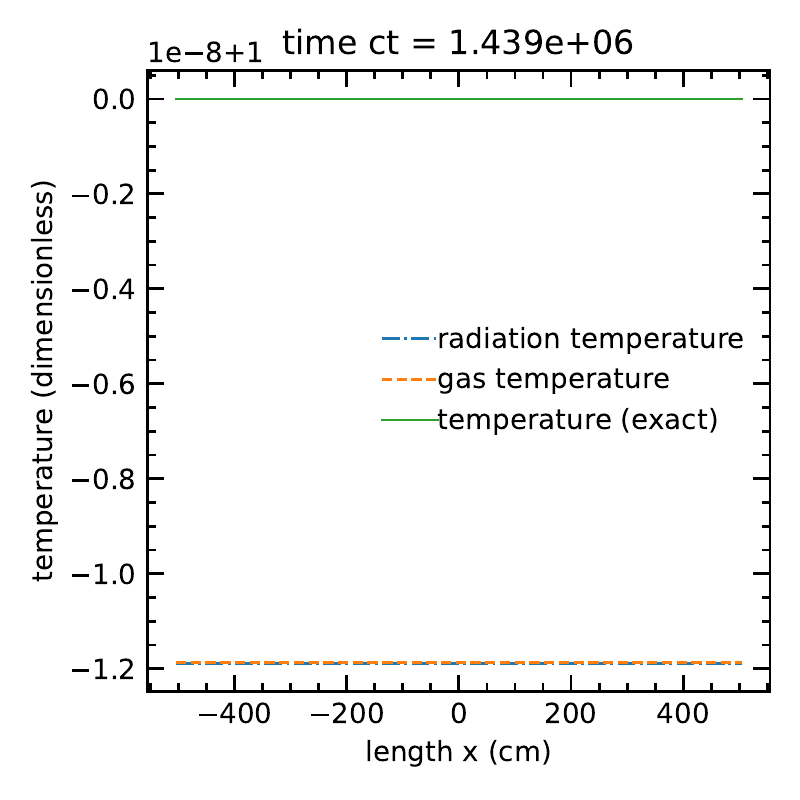 The radiation temperature and matter temperatures, along with the exact solution.
The radiation temperature and matter temperatures, along with the exact solution.
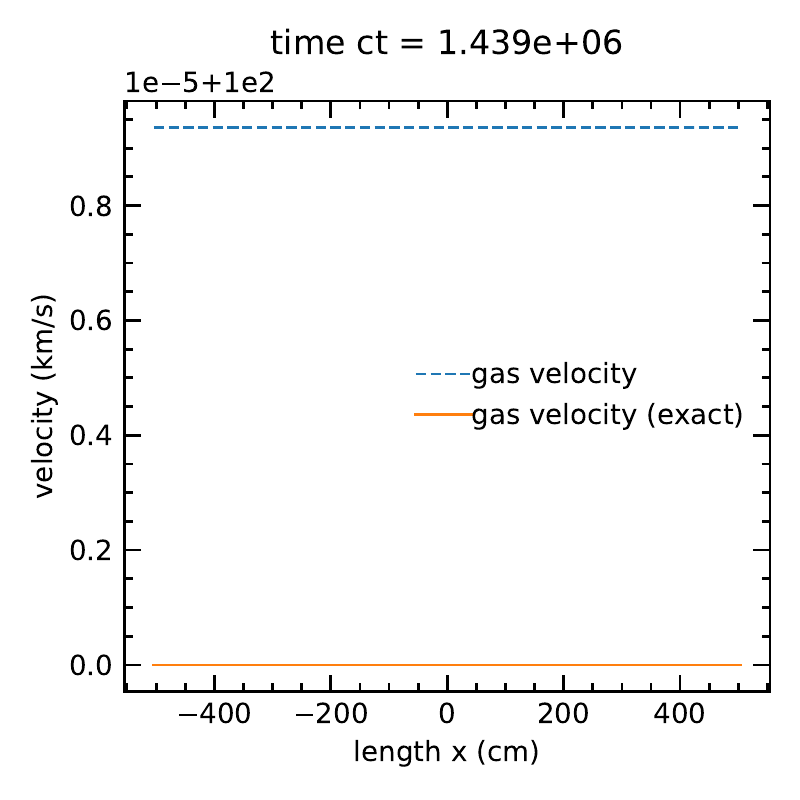 The matter velocity, along with the exact solution.
The matter velocity, along with the exact solution.
Physics
In the transport equation, both the radiation energy and flux are unchanged because the radiation flux and pressure are uniform. In the matter-radiation exchange step, the source term is zero since the radiation and matter are in equilibrium. Finally, the flux is updated following
With \(F_{r}^{(t)} = 4 v E_{r}^{(t)} / 3\), and \(\kappa_P=\kappa_R=\kappa\), we have
Therefore, \(F_r\) remains constant. This demonstrates that the code is invariant under Lorentz transformation.
We can also show that, with the \(O(\beta \tau)\) terms in the matter-radiation exchange step, the space-like component of the radiation four-force vanishes:
Advecting radiation pulse test
This test demonstrates the code’s ability to deal with the relativistic correction source terms that arise from the mixed frame formulation of the RHD moment equations, in a fully-coupled RHD problem. The problems involve the advection of the a pulse of radiation energy in an optically thick (\(\tau \gg 1\)) gas in both static (\(\beta \tau \ll 1\)) and dynamic (\(\beta \tau \gg 1\)) diffusion regimes, with a uniform background flow velocity (Krumholz et al., 2007).
Parameters
Initial condition of the problem in static diffusion regime:
The simulation is run till \(t_{\rm end} = 2 w/v = 4.8 \times 10^{-5} ~{\rm s}\).
Initial condition of the problem in dynamic diffusion regime: same parameters as in the static diffusion regime except
Results
Static diffusion regime:
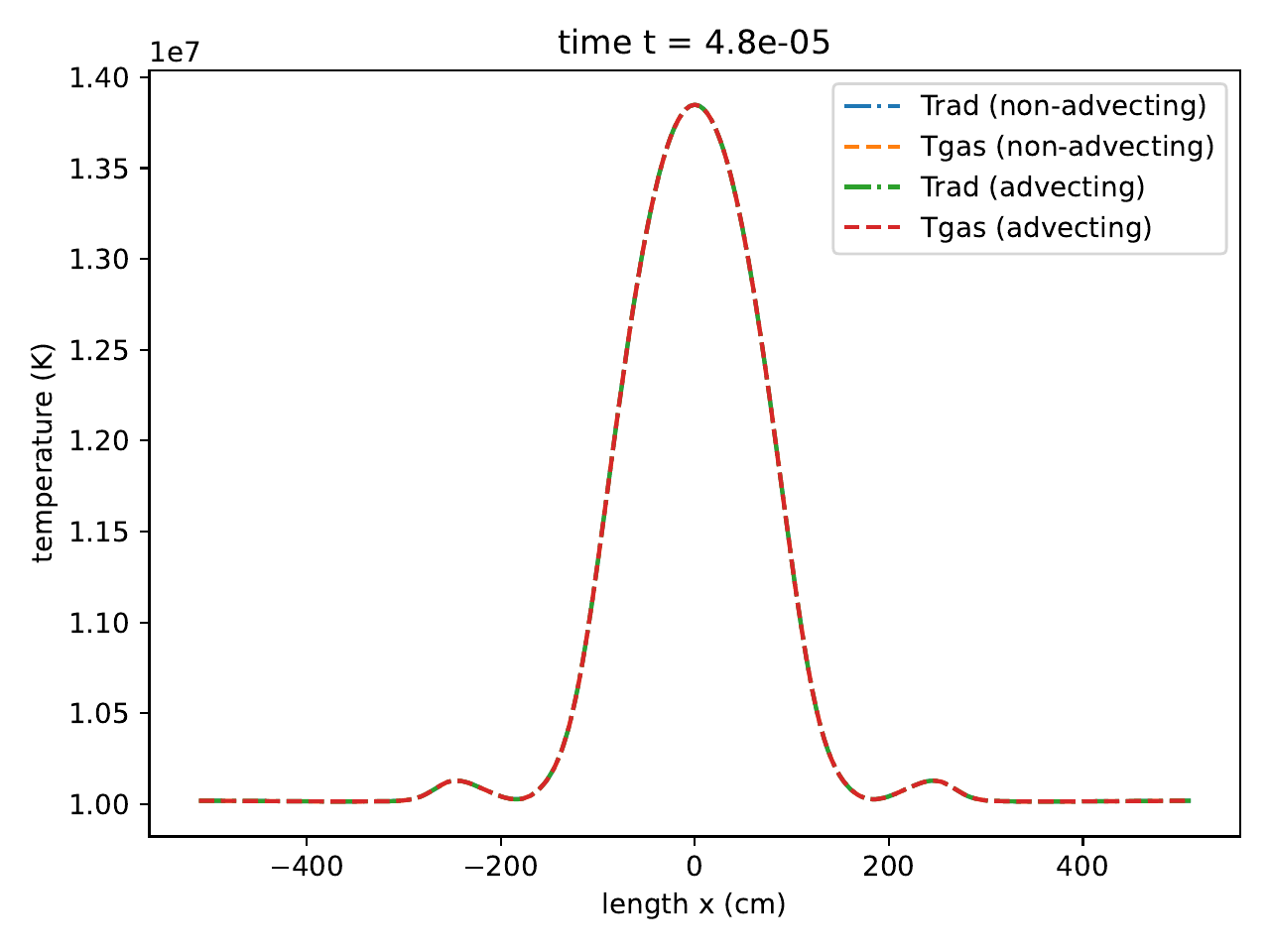 radhydro_pulse_temperature-static-diffusion
radhydro_pulse_temperature-static-diffusion
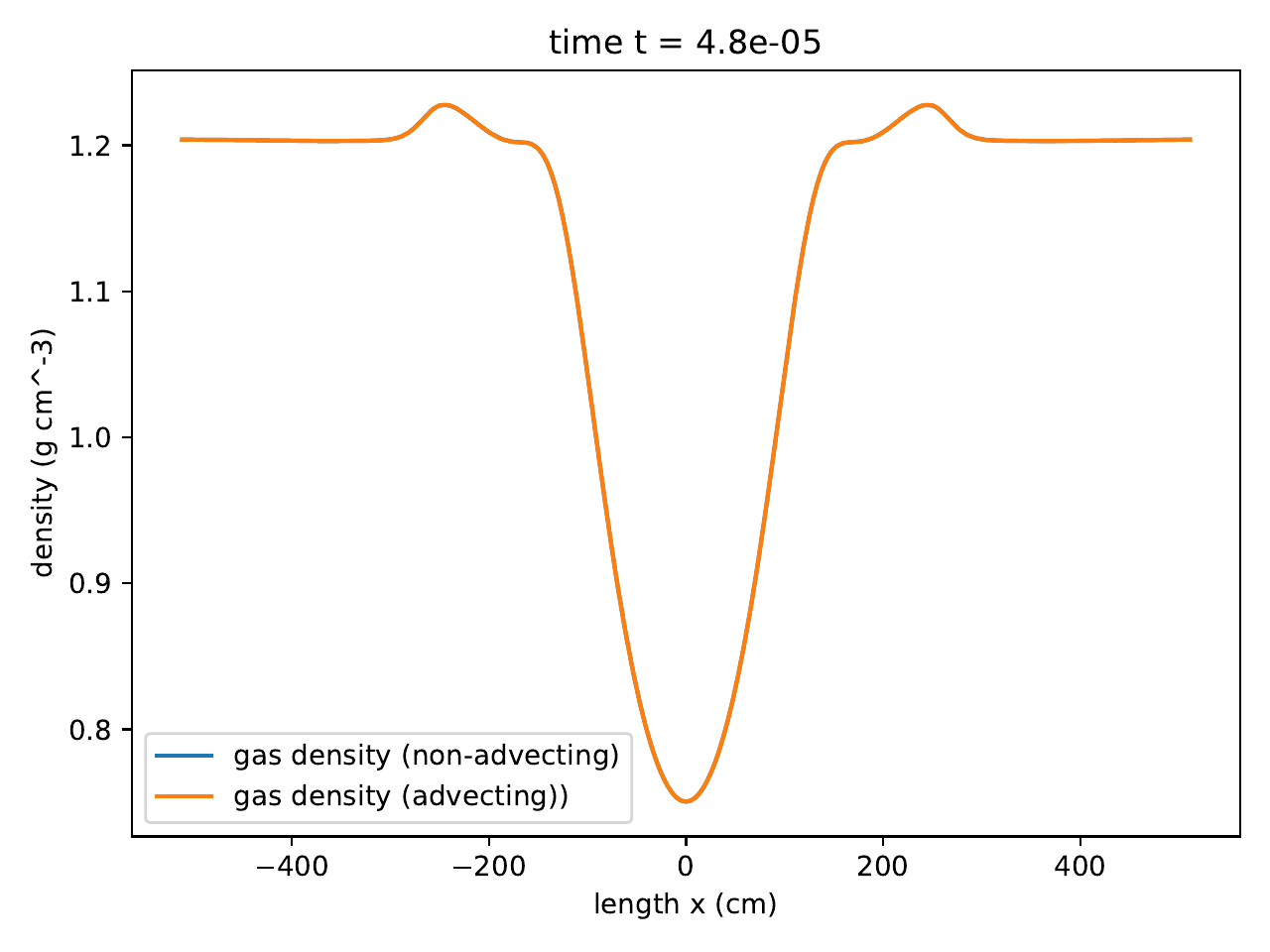 radhydro_pulse_density-static-diffusion
radhydro_pulse_density-static-diffusion
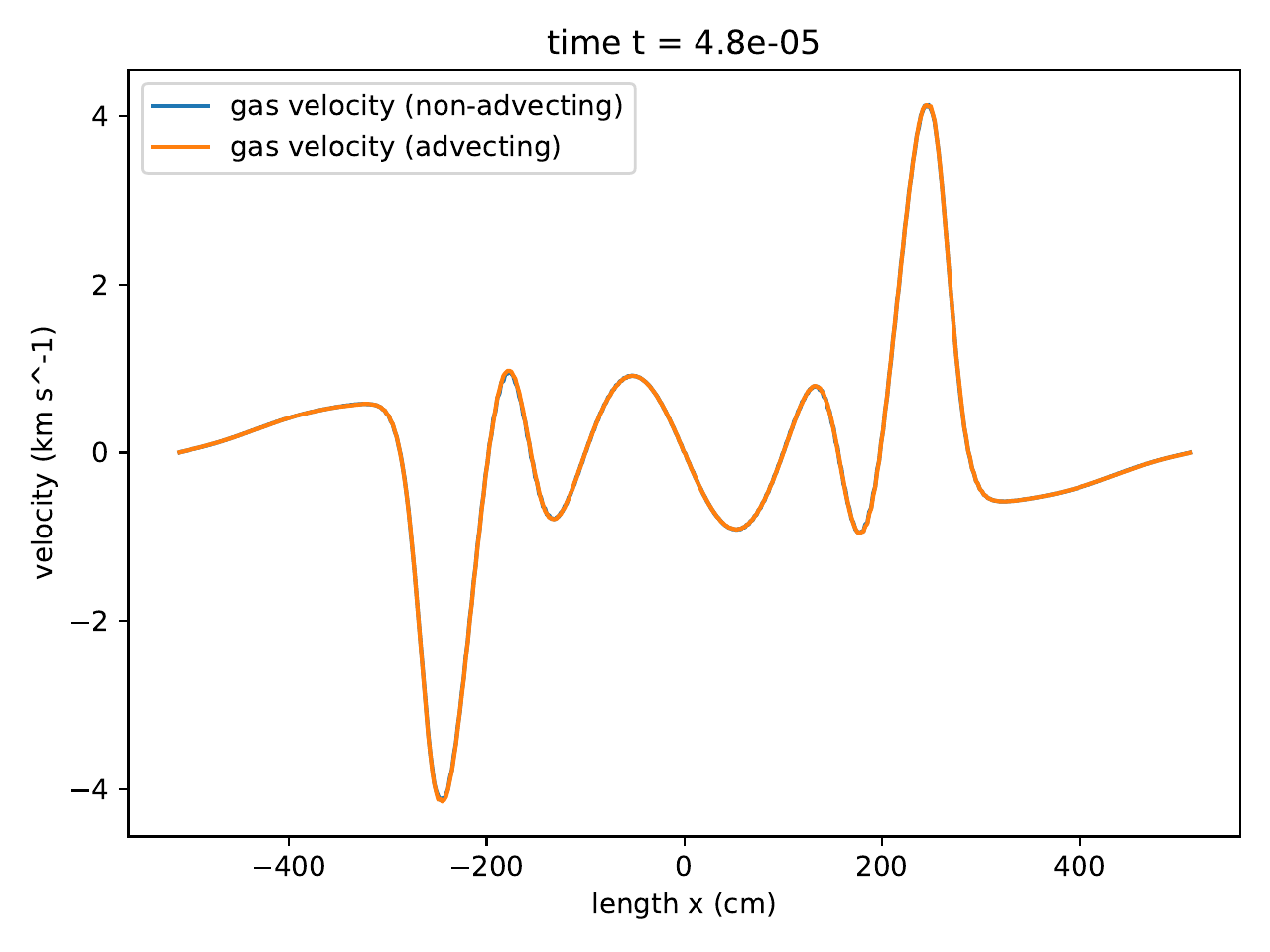 radhydro_pulse_velocity-static-diffusion
radhydro_pulse_velocity-static-diffusion
Dynamic diffusion regime:
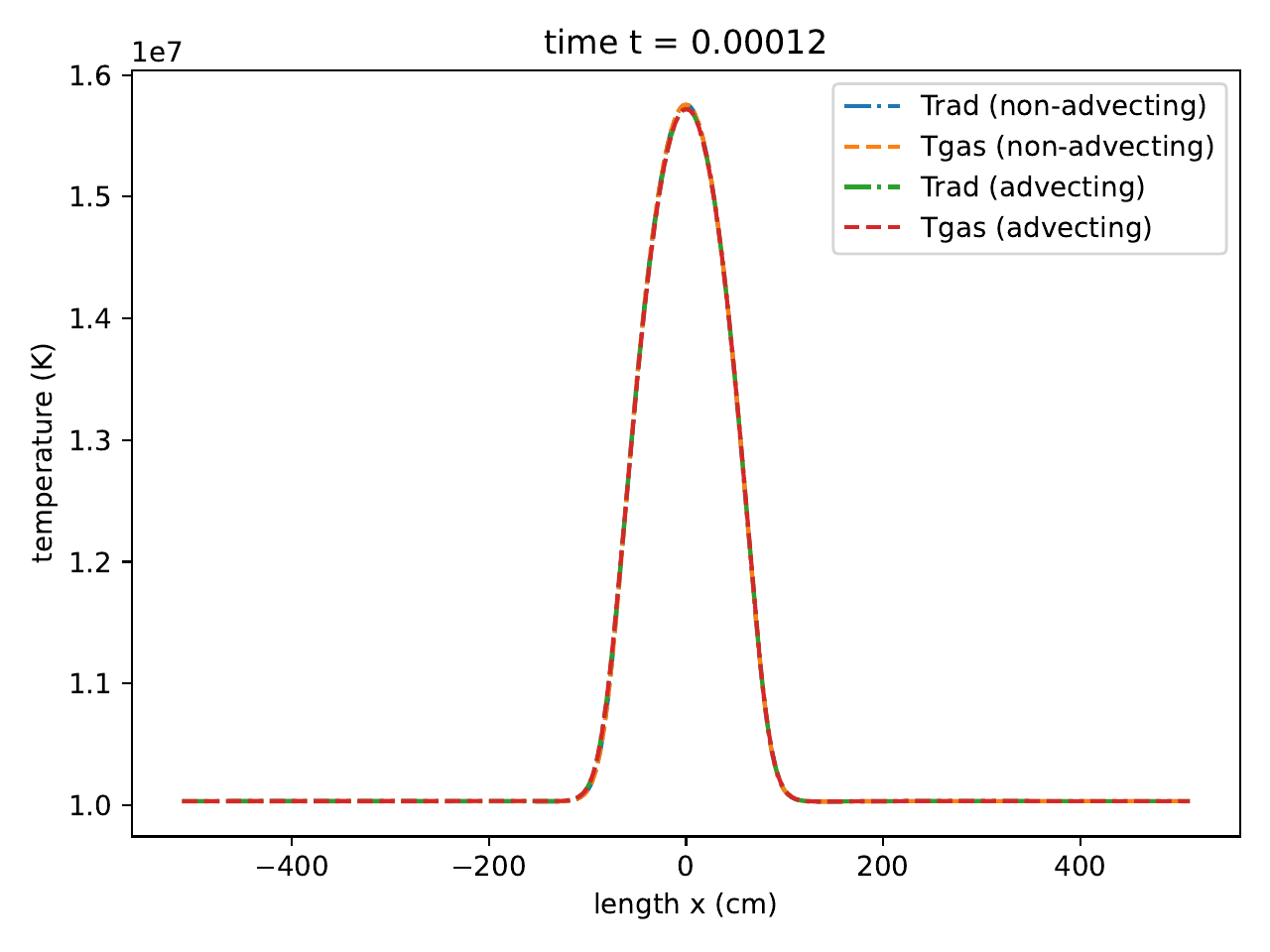 radhydro_pulse_temperature-dynamic-diffusion
radhydro_pulse_temperature-dynamic-diffusion
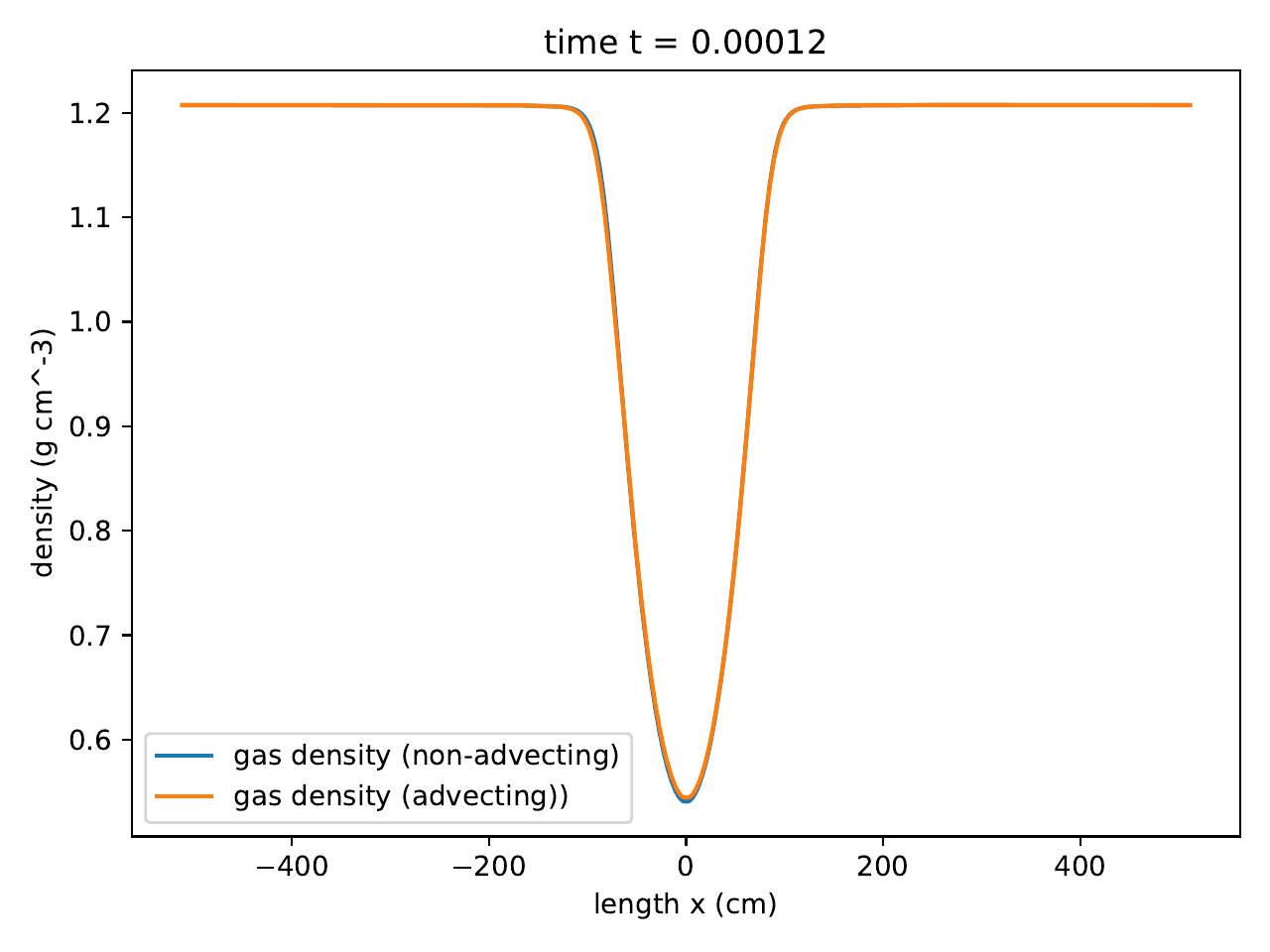 radhydro_pulse_density-dynamic-diffusion
radhydro_pulse_density-dynamic-diffusion
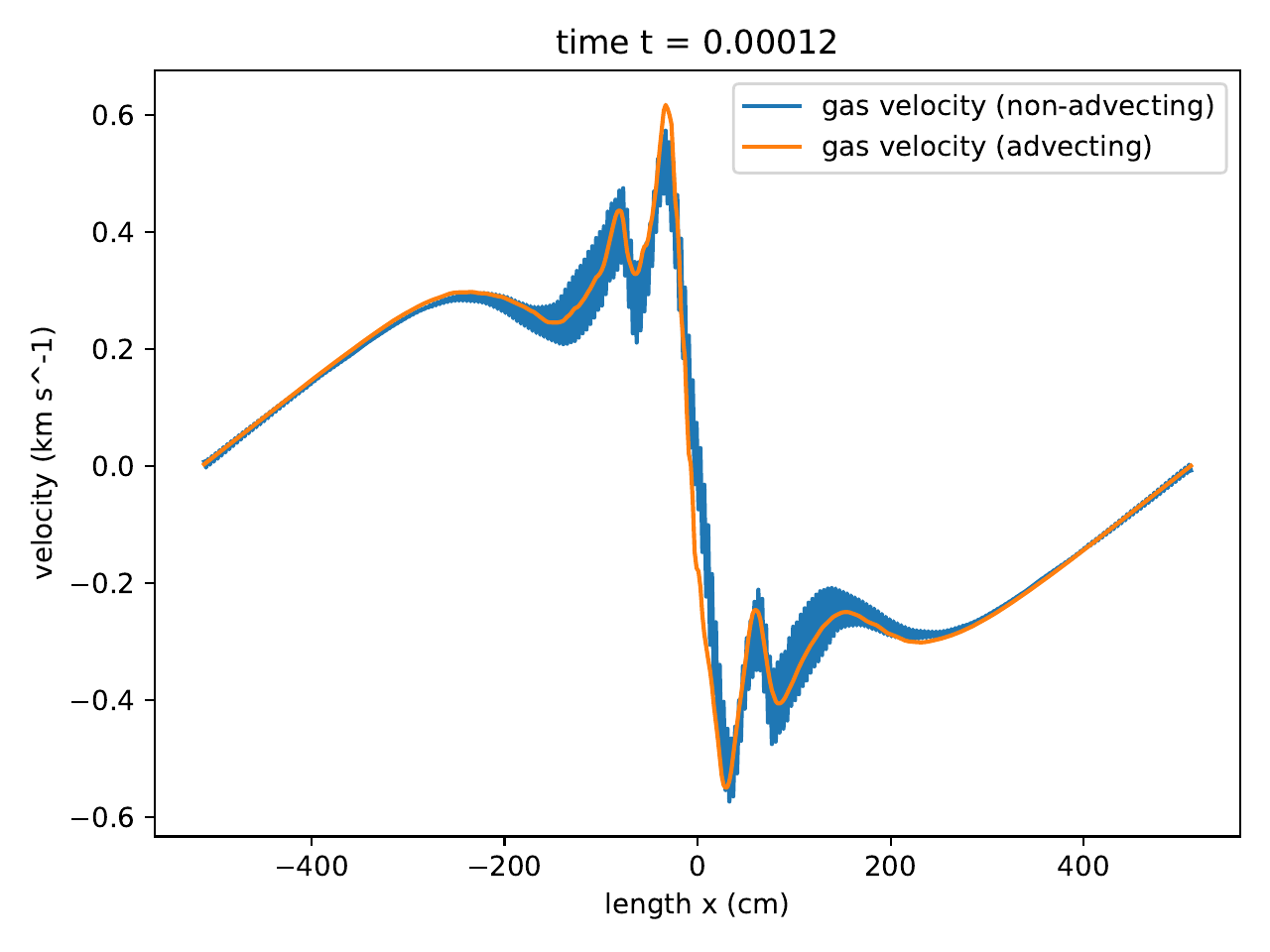 radhydro_pulse_velocity-dynamic-diffusion
radhydro_pulse_velocity-dynamic-diffusion
Runtime parameters
This document lists all of the runtime parameters in Quokka that are set using the AMReX ParmParse object. Users can set these via the input file or by using command-line arguments.
General
These parameters are read in the AMRSimulation<problem_t>::readParameters() function in src/simulation.hpp or src/main.cpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| max_timesteps | Integer | INT_MAX | The maximum number of time steps for the simulation. |
| cfl | Float | 0.3 | Sets the CFL number for the simulation. |
| amr_interpolation_method | Integer | 1 | Selects the method used to interpolate from coarse to fine AMR levels. 0: piecewise constant, 1: linear. Except for debugging, this should not be changed. |
| stop_time | Float | 1.0 | The simulation time at which to stop evolving the simulation. |
| plotfile_interval | Integer | -1 (Disabled) | The number of coarse timesteps between plotfile outputs. |
| plottime_interval | Float | -1.0 (Disabled) | The time interval (in simulated time) between plotfile outputs. |
| skip_initial_plotfile | Boolean (0/1) | 0 (False) | Skip writing the initial plotfile at t=0. |
| statistics_interval | Integer | -1 (Disabled) | The number of coarse timesteps between statistics outputs. |
| checkpoint_interval | Integer | -1 (Disabled) | The number of coarse timesteps between checkpoint outputs. |
| checkpointtime_interval | Float | -1.0 (Disabled) | The time interval (in simulated time) between checkpoint outputs. |
| do_reflux | Boolean (0/1) | 1 (Enabled) | This turns on refluxing at coarse-fine boundaries (1) or turns it off (0). Except for debugging, this should always be on when AMR is used. |
| do_tracers | Boolean (0/1) | 0 (Disabled) | This turns on tracer particles. They are initialized one-per-cell and they follow the fluid velocity. |
| suppress_output | Boolean (0/1) | 0 (Disabled) | If set to 1, this disables output to stdout while the simulation is running. |
| derived_vars | String list | Empty | A list of the names of derived variables that should be included in plotfile outputs. |
| regrid_interval | Integer | 2 | The number of timesteps between AMR regridding. |
| density_floor | Float | 0.0 | The minimum density value allowed in the simulation. Enforced through EnforceLimits. |
| density_floor_expr | String | Empty | Optional AMReX parser expression for a spatially varying density floor. Variables: x, y, z, base_density_floor. When set, this overrides the constant floor. |
| heating_rate_external | String | Empty | Optional AMReX parser expression for external heating rate per H atom (erg/s/H). Variables: time, dt. Effective when cooling is enabled. |
| debug_density_floor_plot | Boolean (0/1) | 0 (Disabled) | If set to 1, adds a derived field density_floor_dbg to plotfiles to visualize the spatially varying density floor. |
| temperature_floor | Float | 0.0 | The minimum temperature value allowed in the simulation. Enforced through EnforceLimits. |
| max_walltime | String | 0 (Unlimited) | The maximum walltime for the simulation in the format DD:HH:SS (days/hours/seconds). After 90% of this walltime elapses, the simulation will automatically stop and exit. |
| dt_cutoff | Float | 0.0 (Disabled) | Timestep drop detector threshold. If the timestep drops below dt_cutoff * current_time, the simulation aborts with an error message. This helps detect numerical instabilities early. |
| constant_dt | Float | 0.0 (Disabled) | Optional constant timestep. If set, forces the timestamp to be this value. |
| initial_dt | Float | Unlimited | Optional initial timestep. |
| max_dt | Float | Unlimited | Optional maximum timestep. |
| init_shrink | Float | 1.0 | Factor to shrink the initial timestep by. |
| show_performance_hints | Boolean (0/1) | 1 (Enabled) | If set to 1, prints performance hints. |
| signal_speed_abort | Float | -1.0 (Disabled) | If value > 0 and the max signal speed exceeds this value, the simulation aborts. |
| particle_speed_abort | Float | -1.0 (Disabled) | If value > 0 and the max particle speed exceeds this value, the simulation aborts. |
| sfh_interval | Integer | -1 (Disabled) | Interval for updating/writing star formation history. |
| sfh_time_interval | Float | -1.0 (Disabled) | Time interval for updating/writing star formation history. |
| particle_cfl | Float | 0.5 | Sets the CFL number for particle advection. This is independent of the hydro CFL number. |
| plotfile_prefix | String | "plt" | The prefix for plotfile output filenames. |
| checkpoint_prefix | String | "chk" | The prefix for checkpoint output filenames. |
| do_subcycle | Boolean (0/1) | 1 (Enabled) | This turns on subcycling at coarse-fine boundaries (1) or turns it off (0). |
| poisson_supercycle_interval | Integer | 1 | The number of coarse timesteps between Poisson supercycle operations. |
| poisson_reltol | Float | 1.0e-5 | Relative tolerance for the Poisson solver convergence. |
| poisson_abstol | Float | 1.0e-5 | Absolute tolerance for the Poisson solver convergence (scaled by minimum RHS value). |
| print_cycle_timing | Boolean (0/1) | 0 (Disabled) | If set to 1, prints per-cycle timing information. |
| restartfile | String | Empty | The path to a checkpoint file from which to restart the simulation. |
| amr.plot_nfiles | Integer | -1 (Auto) | Maximum number of binary files per multifab for plotfiles. Controls parallel I/O chunking. |
| amr.checkpoint_nfiles | Integer | -1 (Auto) | Maximum number of binary files per multifab for checkpoints. Controls parallel I/O chunking. |
| quokka.bc | String list | Required | Boundary conditions for the domain faces. Must be a list of 3 strings (e.g., periodic periodic reflecting). Overrides geometry.is_periodic. |
Hydrodynamics
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| hydro.low_level_debugging_output | Boolean (0/1) | 0 (Disabled) | If set to 1, turns on low-level debugging output for each RK stage. Warning: this writes an enormous volume of data to disk! This should only be used for debugging. |
| hydro.rk_integrator_order | Integer | 2 (RK2-SSP) | Determines the order of the RK integrator used. Can be set to 1 (Forward Euler) or 2 (RK2-SSP, also known as Heun’s method). This should only be changed for debugging. |
| hydro.reconstruction_order | Integer | 3 (PPM) | Determines the order of spatial reconstruction algorithm used. Can be set to 1 (piecewise constant), 2 (piecewise linear; PLM), or 3 (piecewise parabolic; PPM). |
| hydro.plm_limiter | String | "sweby" | Selects the slope limiter for PLM reconstruction. Options: minmod, sweby, or mc. Only used when hydro.reconstruction_order = 2. |
| hydro.use_dual_energy | Boolean (0/1) | 1 (Enabled) | If set to 1, the code evolves an auxiliary internal energy variable in order to correctly evolve high-mach flows. This should only be disabled (0) for debugging. |
| hydro.abort_on_fofc_failure | Boolean (0/1) | 1 (Enabled) | If set to 1, the code aborts when first-order flux correction fails to yield a physical state (positive density and pressure). This should only be disabled (0) for debugging. |
| hydro.artificial_viscosity_coefficient | Float | 0.0 | This is the linear artificial viscosity coefficient used in the artificial viscosity term added to the flux. This is the same parameter as defined in the original PPM paper. |
Radiation
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| radiation.reconstruction_order | Integer | 3 (PPM) | Determines the order of spatial reconstruction algorithm used. Can be set to 1 (piecewise constant), 2 (piecewise linear; PLM), or 3 (piecewise parabolic; PPM). |
| radiation.cfl | Float | 0.3 | Sets the CFL number for the radiation advance. This is independent of the hydro CFL number. |
| radiation.dust_gas_interaction_coeff | Float | 2.5e-34 | Coefficient for dust-gas interaction in radiation calculations. |
| radiation.print_iteration_counts | Boolean (0/1) | 0 (Disabled) | If set to 1, prints radiation iteration counts for debugging. |
| radiation.iteration_tolerance | Float | 1e-11 | Tolerance for the Newton-Raphson iteration residuals. |
| radiation.iteration_tolerance_rel | Float | -1.0 (Disabled) | Tolerance for the relative change between two consecutive Newton-Raphson iterations. |
MHD
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| mhd.emf_computing_scheme | String | "FelkerStone2017" | Determines the method used to compute the EMF at edges. Can be set to FelkerStone2017, Balsara2025, or Quokka2026. |
| mhd.emf_averaging_scheme | String | "LondrilloDelZanna2004" | Determines the method used to average EMF at edges. Can be set to LondrilloDelZanna2004 or Balsara2025. |
| mhd.emf_reconstruction_order | Integer | 5 (xPPM) | Determines the order of spatial reconstruction algorithm used for EMF computation. Can be set to 1 (piecewise constant), 2 (piecewise linear; PLM), 3 (piecewise parabolic; PPM), or 5 (extrema-preserving xPPM). |
| mhd.plm_limiter | String | "sweby" | Selects the slope limiter for PLM reconstruction in EMF calculations. Options: minmod, sweby, or mc. Only used when mhd.emf_reconstruction_order = 2. |
| mhd.project_initial_b_field | Boolean (0/1) | 0 (Disabled) | If set to 1, projects the initial magnetic field to be divergence-free. |
| mhd.update_initial_b_energy | Boolean (0/1) | 1 (Enabled) | If set to 1, updates the initial magnetic energy after projection. |
Optically-thin radiative cooling
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| cooling.enabled | Boolean (0/1) | 0 (Disabled) | If set to 1, turns on optically-thin radiative cooling as a Strang-split source term. |
| cooling.cooling_table_type | String | "resampled" | Specifies the type of cooling table to use. The only supported option is “resampled”. |
| cooling.read_tables_even_if_disabled | Boolean (0/1) | 0 (Disabled) | If set to 1, reads the cooling tables even if the cooling module is disabled. |
| cooling.hdf5_data_file | String | Required if cooling.enabled=1 | The path to the cooling tables in HDF5 format. We recommend using extern/cooling/CloudyData_UVB=HM2012_resampled.h5 for ISM at solar metallicity. |
Chemistry
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| chemistry.enabled | Boolean (0/1) | 0 (Disabled) | If set to 1, turns on chemistry as a Strang-split source term. |
| chemistry.max_density_allowed | Float | 1.0e300 | Maximum density value for which chemistry calculations are accurate. Chemistry is not performed for cells with densities above this threshold. |
| chemistry.min_density_allowed | Float | Smallest positive Value | Minimum density value for which chemistry calculations are performed. Chemistry is not performed for cells with densities below this threshold. |
Dust
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| dust.enable_iter_stoptime | Boolean (0/1) | 0 (Disabled) | If set to 1, enables iterative dust stopping time calculation. |
| dust.omega | Float | 1.0 | Controls the level of frictional heating, with omega = 0 turning it off and omega = 1 depositing all dissipation into the gas. |
| dust.print_iteration_counts | Boolean (0/1) | 0 (Disabled) | If set to 1, prints dust drag iteration counts for debugging. |
| dust.density_floor | Float | 0.0 | The minimum dust density value allowed in the simulation. Enforced through EnforceLimits. |
Particles
These parameters are read in the particleParmParse() function in src/particles/particle_types.hpp and readParmParse() in src/simulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| particles.disable_SN_feedback | Boolean (0/1) | 0 | If set to 1, disables SN feedback when a particle evolves from SNProgenitor to SNRemnant. |
| particles.sink_particle_use_uniform_kernel | Boolean (0/1) | 0 (Disabled) | If set to 1, uses uniform accretion kernel in a (7 dx)^3 box for sink particles. |
| particles.SN_scheme | String | SN_thermal_or_thermal_momentum | Scheme for SN feedback. Options: SN_thermal_only, SN_thermal_or_thermal_momentum, SN_thermal_kinetic_or_thermal_momentum, SN_pure_kinetic_or_thermal_momentum. |
| particles.SN_p_term_Msunkmps | Float | 2.8e5 | Terminal momentum of the supernova remnant in units of \(M_\odot,\mathrm{km,s}^{-1}\). The shell-formation mass \(M_\mathrm{sf}\) is scaled as \((p/p_\mathrm{canonical})^2\) so that the kinetic energy \(p^2/(2M_\mathrm{sf})\) is preserved. |
| particles.eps_ff | Float | 0.01 | Star formation efficiency parameter. |
| particles.verbose | Boolean (0/1) | 0 | Verbosity level for particle operations. Higher values provide more detailed output. |
| particles.param1 | Float | -1.0 | Placeholder parameter for particles (used in gravity_3d.cpp tests). |
| particles.param2 | Float | -1.0 | Placeholder parameter for particles (used in gravity_3d.cpp tests). |
| particles.disable_particle_drift | Boolean (0/1) | 0 | If set to 1, disables particle drift. |
| particles.stellar_velocity_limit | Float | 1.0e8 | Maximum velocity limit for stellar particles in cm/s. The code will abort if stellar velocity exceeds this limit. |
| particles.reproducibility_roundoff_redundancy | Integer | 20 | Number of bits to remove from the significand for reproducibility. |
| particles.use_luminosity_table | Boolean (0/1) | 1 (Enabled) | If set to 1, uses a luminosity table for particles. |
| particles.rad_table | String | Required if particles.use_luminosity_table=1 and Physics_Traits<problem_t>::is_radiation_enabled=true | Path to the radiation luminosity table. |
| particles.rad_table_output_spacing | Integer | 0 (fast_log) | Output spacing for radiation table. |
| particles.split_particles_on_restart_refine | Boolean (0/1) | 1 (Enabled) | Whether to split particles when restarting with refinement. |
Turbulence
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp. Details about the turbulence driving implementation can be found here.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| turbulence.enabled | Boolean (0/1) | 0 (Disabled) | If set to 1, enables turbulence driving using chfeder’s turbulence driving module. |
| turbulence.length | Float | Required if turbulence.enabled=1 | Length of turbulent driving box. Can have 1 value or a comma separated list for each component. |
| turbulence.target_vdisp | Float | Required if turbulence.enabled=1 | Target turbulent velocity dispersion. |
| turbulence.ampl_factor | Float | Required if turbulence.enabled=1 | Amplitude adjust factor for forcing field. Can have 1 value or a comma separated list for each component. |
| turbulence.ampl_auto_adjust | Boolean (0/1) | 0 (Disabled) | If set to 1, enables automatic amplitude adjustment. |
| turbulence.k_driv | Float | Required if turbulence.enabled=1 | Characteristic driving scale in units of \(2\pi/length\). This sets the autocorrelation timescale via tau = (length/target_vdisp)/k_driv. |
| turbulence.k_min | Float | Required if turbulence.enabled=1 | Minimum driving wavenumber in units of \(2\pi/length\). |
| turbulence.k_max | Float | Required if turbulence.enabled=1 | Maximum driving wavenumber in units of \(2\pi/length\). |
| turbulence.sol_weight | Float | Required if turbulence.enabled=1 | Solenoidal weight. Can be 0.0 (compressive driving), 1.0 (solenoidal driving) or 0.5 (natural mixture). |
| turbulence.spect_form | Integer | Required if turbulence.enabled=1 | Spectral form of the driving amplitude. Can be 0 (band, rectangle, constant), 1 (paraboloid) or 2 (power law). |
| turbulence.power_law_exp | Float | Required if turbulence.spect_form=2 | If spect_form = 2, this sets the spectral power-law exponent (e.g., -5/3: Kolmogorov; -2: Burgers). |
| turbulence.angles_exp | Float | Required if turbulence.spect_form=2 | If spect_form = 2, this sets the number of modes (angles) in k-shell such that it increases as \(k^angles_exp\). |
| turbulence.stop_time | Float | Optional if turbulence.enabled=1 | Time at which to stop turbulence driving. Default is max, i.e. never stop. |
| turbulence.random_seed | Integer | Required if turbulence.enabled=1 | Random number seed for driving sequence. |
| turbulence.nsteps_per_t_turb | Integer | Required if turbulence.enabled=1 | Number of turbulence driving pattern updates per turnover time. |
Photoelectric Heating
These parameters are read in the QuokkaSimulation<problem_t>::readParmParse() function in src/QuokkaSimulation.hpp.
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| use_sfh_based_pe_heating | Boolean (0/1) | 0 (Disabled) | If set to 1, enables photoelectric heating based on star formation history. |
| sfh_to_pe_heating_table | String | Required if use_sfh_based_pe_heating = 1 | Path to the table converting star formation history to photoelectric heating. |
| sf_area_kpc2 | Float | Required if use_sfh_based_pe_heating = 1 and const_sfr_Msun_per_year_per_kpc2 < 0 | Area of the star formation region in kpc^2. |
| const_sfr_Msun_per_year_per_kpc2 | Float | -1.0 | Constant star formation rate in Msun/year/kpc^2. If non-negative, overrides the simulation SFR. |
In-situ analysis
In-situ analysis refers to analyzing simulations as they are running using Quokka’s built-in runtime diagnostics.
Diagnostics
Most of Quokka’s diagnostics are adapted from the implementation included in the Pele suite of AMReX-based combustion codes. (See the documentation for PeleLMeX diagnostics for an explanation of the original implementation.)
There are three built-in diagnostics that can be configured to output at periodic intervals while the simulation is running:
- axis-aligned 2D projections
- axis-aligned 2D slices, and
- N-dimensional probability distribution functions (PDFs).
2D Projections
This diagnostic outputs 2D axis-aligned projections as AMReX plotfiles prefixed with proj and is configured using the ProjectionPlot diagnostic type. Only one direction is supported per diagnostic; configure multiple diagnostics to output multiple directions.
Note
Filters are not supported for ProjectionPlot diagnostics and will be ignored with a warning.
Note
Projection outputs are line integrals along the projection axis, using code length units. Each cell value is multiplied by
dx[dir]and summed, so the output units are (field units) × length.Examples:
nH(cm^-3) → projectednH(cm^-2) column density;pressure(K cm^-3) → projected pressure (K cm^-2). Avoid projecting per-cell extensive quantities likemass = rho * dVunless you intentionally want an extra length factor.
AMR Structure Preservation: As of recent updates, 2D projections now preserve the full AMR structure from the 3D simulation, maintaining higher resolution where needed instead of averaging down to level 0. This provides better detail but may result in larger output files.
This diagnostic is configured entirely with runtime parameters. Use field_names to select which plotfile/derived variables to project along the selected direction.
Example input file configuration (defaults shown as inline comments):
quokka.diagnostics = proj_x proj_z # enable two diagnostics with independent settings
quokka.proj_x.type = ProjectionPlot # required: selects 2D projections
quokka.proj_x.file = proj/x/plt # output path prefix (subdir + "pltNNNNNNN")
quokka.proj_x.int = 200 # output cadence (coarse steps)
quokka.proj_x.normal = 0 # 0==x, 1==y, 2==z (default: 0)
quokka.proj_x.field_names = nH temperature # fields to project (plotfile/derived vars)
quokka.proj_z.type = ProjectionPlot
quokka.proj_z.file = proj/z/plt
quokka.proj_z.int = 200
quokka.proj_z.normal = 2
quokka.proj_z.field_names = nH
2D Slices
Note
This is based on the DiagFramePlane diagnostic from PelePhysics, and the same runtime parameters should apply here without modification. The output format is also the same as that produced by the Pele codes.
This outputs 2D slices of the simulation as AMReX plotfiles that can be further examined using, e.g., VisIt or yt.
Example input file configuration:
quokka.diagnostics = slice_z # Space-separated name(s) of diagnostics (arbitrary)
quokka.slice_z.type = DiagFramePlane # Diagnostic type (others may be added in the future)
quokka.slice_z.file = slicez_plt # Output file prefix (should end in "plt")
quokka.slice_z.normal = 2 # Plane normal (0 == x, 1 == y, 2 == z)
quokka.slice_z.center = 2.4688e20 # Coordinate in the normal direction
quokka.slice_z.int = 10 # Output interval (in number of coarse steps)
# The problem must output these derived variable(s)
derived_vars = temperature
# List of variables to include in output
quokka.slice_z.field_names = gasDensity gasInternalEnergy temperature
Histograms/PDFs
Note
This is based on the DiagPDF diagnostic from PelePhysics, but significant changes have been made to both the runtime parameters and the output format in order to support N-dimensional histograms, log-spaced binning, and histogramming by mass.
This adds histogram outputs (as fixed-width text files) at fixed timestep intervals as the simulation evolves. The quantity accumulated in each bin is the total mass, volume, or cell count summed over all cells not covered by refined grids over all AMR levels. If unspecified in the input parameters, the default is to accumulate the volume in each bin.
By default, the bins extend over the full range of the data at a given timestep. The range parameter can instead specify the minimum and maximum extent for the bins. Bins can be optionally log-spaced by setting log_spaced_bins = 1.
Normalization of the output is left up to the user.
Example input file configuration:
quokka.hist_temp.type = DiagPDF # Diagnostic type
quokka.hist_temp.file = PDFTempDens # Output file prefix
quokka.hist_temp.int = 10 # Output cadence (in number of coarse steps)
quokka.hist_temp.weight_by = mass # (Optional, default: volume) Accumulate: mass, volume, cell_counts
quokka.hist_temp.var_names = temperature gasDensity # Variable(s) of interest (compute a N-D histogram)
quokka.hist_temp.temperature.nBins = 20 # temperature: Number of bins
quokka.hist_temp.temperature.log_spaced_bins = 1 # temperature: (Optional, default: 0) Use log-spaced bins
quokka.hist_temp.temperature.range = 1e3 1e7 # temperature: (Optional, default: data range) Specify min/max of bins
quokka.hist_temp.gasDensity.nBins = 5 # gasDensity: Number of bins
quokka.hist_temp.gasDensity.log_spaced_bins = 1 # gasDensity: (Optional, default: 0) Use log-spaced bins
quokka.hist_temp.gasDensity.range = 1e-29 1e-23 # gasDensity: (Optional, default: data range) Specify min/max of bins
Filters (based on any variables, not necessary those used for the histogram) can be optionally added:
quokka.hist_temp.filters = dense # (Optional) List of filters
quokka.hist_temp.dense.field_name = gasDensity # Filter field
quokka.hist_temp.dense.value_greater = 1e-25 # Filters: value_greater, value_less, value_inrange
Runtime Derived Fields
Runtime-derived fields are produced by factory-registered providers at runtime and can be consumed by diagnostics (for example, DiagPDF) without adding problem-specific ComputeDerivedVar(...) specializations.
Provider discovery is driven by quokka.<name>.type. Quokka scans the configured quokka.*.type entries, instantiates the ones registered as runtime-derived field providers, and enables only the emitted output fields that appear in derived_vars.
To use a runtime-derived field:
- Configure the provider under
quokka.<group_name>.*. - Add the full names of every desired output field from that provider to
derived_varsso they are available to plotfiles and diagnostics.
Provider configuration parameters:
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| derived_vars | String list | Empty | List only the emitted output field names that should appear in plotfiles and diagnostics. |
| quokka.<name>.type | String | None | Factory type for this provider (for example, DerivedParticleDeposition). |
DerivedParticleDeposition provider parameters:
| Parameter Name | Type | Default | Description |
|---|---|---|---|
| quokka.<name>.particle_types | String list | CIC | Particle types to deposit. Supported values: CIC, CICRad, StochasticStellarPop, Sink, Test. |
| quokka.<name>.deposit_fields | String list | mass | Particle fields to deposit. Supported: mass, birth_mass (only for StochasticStellarPop). |
| quokka.<name>.prefix | String | particle | Output naming prefix. Output names are formed as <prefix>.<ParticleType>.mass_density for mass and <prefix>.<ParticleType>.birth_mass_density for birth_mass. |
| quokka.<name>.mass_min | Real | -inf | Optional lower bound on particle mass for deposition. Particles with mass < mass_min are excluded. |
| quokka.<name>.mass_max | Real | +inf | Optional upper bound on particle mass for deposition. Particles with mass > mass_max are excluded. |
| quokka.<name>.t_age | Real | unset | Optional age threshold. When set, only particles with (tNew[0] - birth_time) <= t_age are deposited. |
| quokka.<name>.normalization_expr | String | unset | Optional AMReX parser expression for a multiplicative normalization constant applied after deposition. |
Notes:
- Only emitted outputs listed in
derived_varsare added to plotfiles and made available to diagnostics. - Provider group names are not valid
derived_varsentries. - These fields can be consumed by diagnostics by name.
- Output name collisions with other derived fields are rejected at startup.
t_ageis only supported for particle types that includebirth_time(CICRad,StochasticStellarPop,Test).birth_massis only supported forStochasticStellarPop; using it with other particle types aborts at startup/runtime.normalization_exprmust evaluate to a scalar constant. Parser constants available:Msun,yr,kpc.
Example:
# List only the provider outputs you want to write.
derived_vars = particle.CIC.mass_density particle.Sink.mass_density
# Configure the provider under quokka.<group_name>.*
quokka.partdep.type = DerivedParticleDeposition
quokka.partdep.particle_types = CIC Sink
quokka.partdep.deposit_fields = mass
quokka.partdep.prefix = particle
# Use one of the runtime-derived outputs in a diagnostic.
quokka.diagnostics = slice_z
quokka.slice_z.type = DiagFramePlane
quokka.slice_z.file = slicez_plt
quokka.slice_z.normal = 2
quokka.slice_z.center = 2.4688e20
quokka.slice_z.int = 10
quokka.slice_z.field_names = particle.CIC.mass_density
Postprocessing
There are several ways to post-process the output of Quokka simulations. AMReX PlotfileTools, yt, and VisIt all allow you to analyze the outputs after they are written to disk.
Amrvis-container
Amrvis-container bundles Amrvis in a Docker/Apptainer image with a browser-based X11 frontend. To browse Quokka plotfiles locally (Docker required), run from the Amrvis-container repo:
./launch_amrvis_browser.sh /path/to/plotfiles
The target directory is bind-mounted to /home/vscode/data in the container. The launcher prints a one-time password; open http://localhost:8080, paste the password, and use the xterm window to start amrvis2d or amrvis3d on your plt* directories.
Tip
On SLURM clusters with Apptainer, pull the image once with
apptainer pull amrvis-container.sif docker://ghcr.io/amrex-codes/amrvis-container:main, then use./launch_amrvis_browser_hpc.sh /path/to/plotfileson a compute node and follow the printed SSH tunnel instructions.
AMReX PlotfileTools
These are self-contained C++ programs (included with AMReX in the Tools/Plotfile subdirectory) that will output a 2D slice (axis-aligned), a 1D slice (axis-aligned), or compute a volume integral given an AMReX plotfile. This works as an alternative to yt and VisIt for basic tasks.
- To compute a volume integral, use fvolumesum.
- To compute a 2D slice plot (axis-aligned planes only), use fsnapshot.
- To compute a 1D slice (axis-aligned directions only, with output as ASCII), use fextract.
Other tools:
- fboxinfo prints out the indices of all the Boxes in a plotfile
- fcompare calculates the absolute and relative errors between plotfiles in L-inf norm
- fextrema calculates the minimum and maximum values of all variables in a plotfile
- fnan determines whether there are any NaNs in a plotfile
- ftime prints the simulation time of each plotfile
- fvarnames prints the names of all the variables in a given plotfile
yt
yt can load Quokka AMReX plotfiles through its Quokka frontend, which is now available in the main yt repository. Until a released yt version includes the frontend, install yt from the main branch with the optional Quokka dependencies:
pip install "yt[quokka] @ git+https://github.com/yt-project/yt.git"
After installation, load a plotfile with yt.load("plt00000") or point yt to any other Quokka plotfile directory. The upstream Quokka frontend documentation is available in the yt documentation source, and a tutorial notebook is available at README.ipynb.
Tip
One of the most useful things to do is to convert the data into a uniform-resolution NumPy array with the covering_grid function.
quick_plot script for batch processing
The quick_plot script in scripts/python/ wraps yt for batch visualization of Quokka outputs. It can process multiple snapshots and generate slice or projection plots. For usage details, see the documentation at the top of the script or run quick_plot -h.
yt-studio for web-based visualization
yt-studio provides a web interface and Python API for visualizing Quokka simulation data. Install it with pip install git+https://github.com/chongchonghe/yt-studio.git, then start the web interface with yt-studio and open http://localhost:5173 in your browser. It supports slice plots, projection plots, volume rendering, multiple colormaps, high-resolution export, optional particle overlays, and AMR grid annotations. For scripts and notebooks, use the QuokkaPlotter methods such as slice() and project().
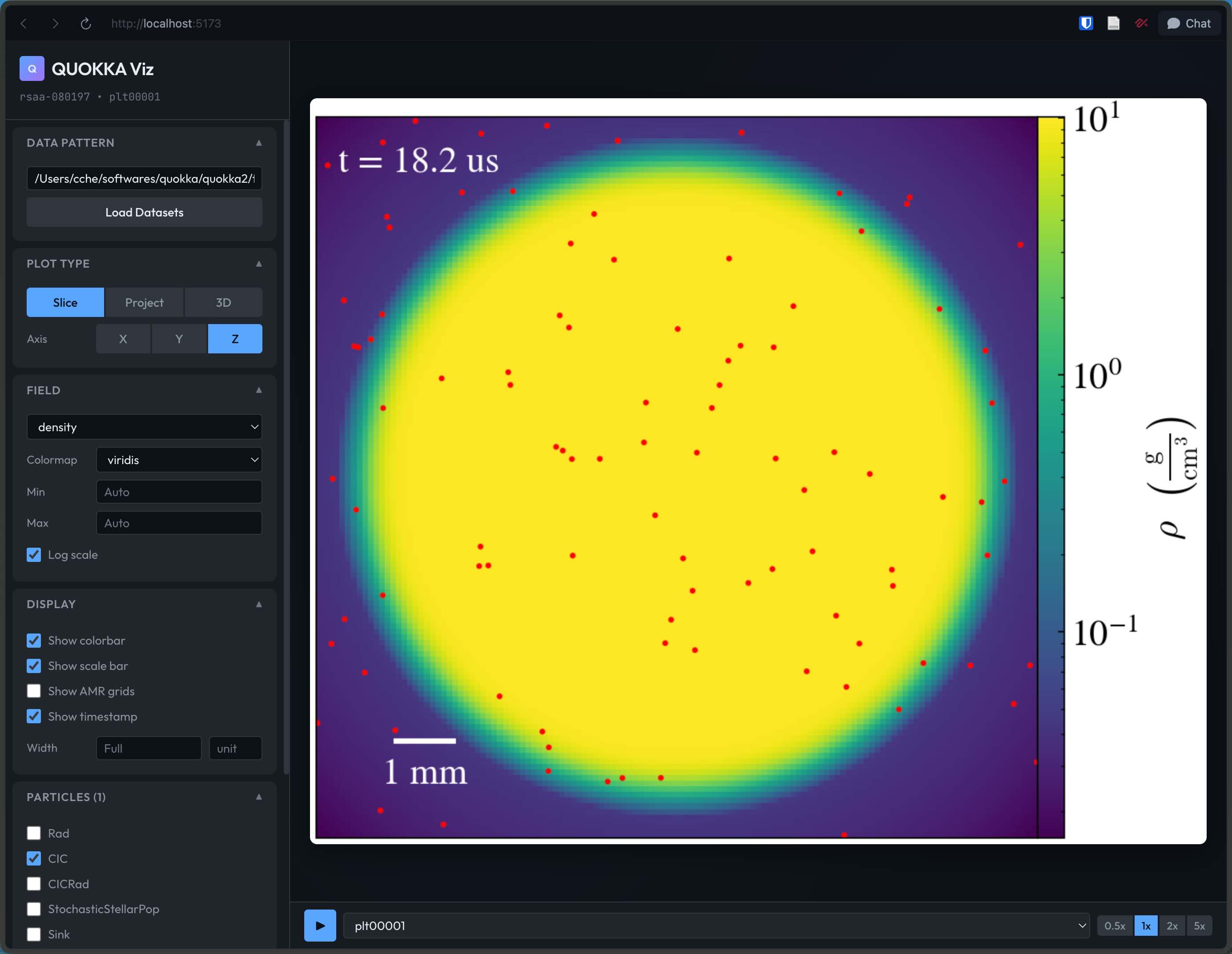
VisIt
VisIt can read cell-centered output variables from AMReX plotfiles. Currently, there is no support for reading either face-centered variables or particles. (However, by default, cell-centered averages of face-centered variables are included in Quokka plotfiles.)
In order to read an individual plotfile, you can select the plt00000/Header file in VisIt’s Open dialog box.
If you want to read a timeseries of plotfiles, you can create a file with a .visit extension that lists the plt*/Header files, one per line, with the following command: :
ls -1 plt*/Header | tee plotfiles.visit
Then select plotfiles.visit in VisIt’s Open dialog box.
Warning
There are rendering bugs with unscaled box dimensions. Slices generally work. However, do not expect volume rendering to work when using, e.g. parsec-size boxes with cgs units.
Debugging simulation instability
Nonlinear stability of systems of PDEs is an unsolved problem. There is no complete, rigorous mathematical theory. There are two concepts, however, that are closely associated with nonlinear stability:
- positivity preservation: This is the property that, given a positive initial density and pressure at timestep \(n\), the density and pressure are positive at timestep \(n+1\). For theoretical background, see Linde & Roe (1997). and Perthame & Shu (1996).
- entropy stability: This is the property that the discretized system of equations obeys the second law of thermodynamics, i.e. the discrete entropy of the simulation must be non-decreasing. There is also a stronger local form, where the entropy variable everywhere obeys an entropy inequality. For theoretical background, see Harten (1983) and Tadmor (1986). (This assumes a convex equation of state.)
If a simulation goes unstable, it is likely due to one of the above properties being violated. It is important to note that standard finite volume reconstruction methods do not guarantee entropy stability (see Fjordholm et al. 2012).
It is also possible that the entropy is nondecreasing, but insufficient entropy is produced for a given shock compared to the amount that should be produced physically. This will cause an unphysical oscillatory solution.
Ways to improve stability
The solution is either to reduce the timestep or add additional dissipation:
- set the initial timestep to be 0.1 or 0.01 of the CFL timestep by setting
sim.initDt_appropriately - lower the CFL number
- It should be in the range 0.1-0.3. If it’s above 0.3, it’s linearly unstable, so it will never work.
- If it’s below 0.1, it’s sufficiently low that the simulation will be very inefficient. If it still doesn’t work, experience indicates that reducing it further usually does not help.
- reduce the order of the spatial reconstruction
- By default PPM reconstruction is used, but PLM (with minmod limiter) can be used instead. It is much more dissipative, and therefore, stable.
- re-try the hydro update with a smaller timestep
- This is necessary because the positivity-preserving timestep may be much smaller than the CFL-limited timestep near the boundary of realizable states (Linde & Roe 1997).
- Quokka will do this automatically, but only up to a maximum hard-coded number of retries.
- If the simulation still fails, this usually indicates a stability problem that will probably not be fixed by further timestep reductions.
- revert to a first-order update in problem cells
- For a sufficiently small timestep, this is provably entropy stable and positivity-preserving (as long as the Riemann solver itself is, which requires robust wavespeeds)
- Quokka reduces to first-order in space and time automatically when the density is negative in a given cell.
- In the future, Quokka could be extended to also revert to first-order based on entropy.
- use wavespeed estimates that are robust for strong shocks
- The eigenvalues of the Roe-average state do not provide correct bounds for very strong shocks.
- If the shocks at the interface travel faster than the wavespeed estimates, there will be insufficient entropy production.
- Doing this requires additional assumptions about the EOS (Miller & Puckett 1996).
- Quokka attempts to do this for ideal gases and as well as materials that can be approximated with a Mie-Gruniesen EOS (see Dukowicz 1985 and Rider 1999).
- No code changes should be required unless you are simulating an exotic material or a condensed matter phase transition (gaseous phase transitions do not cause any issues; see Bethe 1942).
- add artificial viscosity
- This can be helpful because it adds dissipation when shocks are propagating transverse to the interface.
- For sufficient entropy production, it is important that the velocity divergence estimator is based on the cell-average velocities surrounding the interface, not the reconstructed velocities.
- This can be enabled in Quokka with the runtime parameter
hydro.artificial_viscosity_coefficient. A value of0.1is recommended. - This parameter is identical to the artificial viscosity coefficient described in Colella and Woodward 1984.
Floors
As an absolute last resort, one can enable density and/or temperature floors for a simulation using Quokka’s EnforceLimits function.
This may be necessary if the positivity-preserving timestep for a state near vacuum is too small to be feasible. A temperature floor may also be necessary in order to prevent the auxiliary internal energy from becoming negative when there is strong cooling.
Runtime calculator
The runtime calculator is currently a standalone HTML page carried over from the previous documentation site.
- Open the calculator here: Runtime calculator
This wrapper page keeps the calculator reachable from SUMMARY.md within the mdBook site.
Developer onboarding notes
What Quokka is built to do
- Radiation hydrodynamics on AMReX. Quokka targets multi-physics astrophysical simulations using the AMReX adaptive mesh refinement framework, providing two-moment radiation transport, hydrodynamics, and optional MHD capabilities in a single-source C++20 codebase (README.md).
- Modular simulation core. The
AMRSimulationbase class orchestrates time stepping, refinement, and I/O, whileQuokkaSimulationadds domain-specific toggles for radiation, cooling, chemistry, and MHD support that downstream problems can enable selectively (simulation.hpp, QuokkaSimulation.hpp).
Repository tour
src/- Framework layer.
simulation.hppdefinesAMRSimulation, the central driver that manages AMReX state, time stepping, outputs, and particle infrastructure (simulation.hpp). - Physics modules. Hydrodynamics, radiation, MHD, chemistry, and cooling each live in subdirectories such as
hydro/,radiation/, andcooling/, which are wired intoQuokkaSimulationvia thePhysics_Traitsmechanism (QuokkaSimulation.hpp). - Problem drivers. Each scenario in
src/problems/defines aproblem_ttype, customises traits, sets initial conditions, and then instantiatesQuokkaSimulationto run; seesrc/problems/OrszagTang/testOrszagTang.cppfor a representative setup.
- Framework layer.
inputs/- Runtime parameter files that pair with problem drivers; regression entries reference them directly when defining automated tests (regression/quokka-tests.ini).
regression/- The regression harness (
quokka-tests.ini) enumerates long-running GPU test suites, including MPI launch commands, linked data files, and which executables to build (regression/quokka-tests.ini).
- The regression harness (
docs/- Source for the published documentation site (mdBook). The landing page summarises Quokka’s goals and AMReX integration, and additional pages cover workflow diagrams, testing, debugging, and performance topics (site overview, simulation flowchart, test catalog).
Execution flow in practice
- Start-up:
main.cppinitialises AMReX, then callsproblem_main()declared inmain.hppand implemented by each problem driver (main.cpp, main.hpp). - Simulation lifecycle: the flowchart documentation page mirrors the control flow of
AMRSimulation::setInitialConditions,evolve, andcomputeTimestep, showing the nested loops for hydrodynamics stages and radiation subcycling (flowchart overview). - Custom physics: problem-specific traits determine which subsystems
QuokkaSimulationactivates (e.g., MHD, radiation groups, chemistry), and each problem supplies initial conditions (cells and face-centered fields) before callingsim.evolve()(QuokkaSimulation.hpp,testOrszagTang.cpp).
Build, test, and quality checks
- Build locally. Follow the installation guide to clone with submodules, configure with CMake (Ninja or Make), and choose the desired dimensionality and accelerator backend (installation guide).
- Automated tests.
ninja testorctestexercises the bundled problem suite; for full GPU coverage, rely on the regression harness described earlier (installation guide, quokka-tests.ini). - Static analysis. Run
clang-tidymanually or viascripts/tidy.shto match the repository’s CI checks, as documented in the How to Use clang-tidy guide (clang-tidy how-to). - CUDA builds on macOS or Linux. To build and test CUDA functionality locally, run
./scripts/bash/run-cuda-container.sh. This script pulls the appropriate Docker image, launches a container, and performs a CUDA build—useful for catching CUDA-specific issues on your development machine. - Writing and building documentation. The developer is responsible for updating the documentation site. To build and view the documentation site locally:
./scripts/bash/install_mdbook.sh
./scripts/bash/docs_build_and_view.sh
This uses the same mdBook install commands as the GitHub Actions docs jobs.
Then open the printed http://localhost:<port>/ URL in your browser to view the documentation locally.
Where to dive deeper next
- Physics modules. Explore
src/hydro/andsrc/radiation/alongside the corresponding documentation pages (hydro_integrator.md, radiation topics) to understand scheme implementations (QuokkaSimulation.hpp, overview page). - Problem setups. Study more drivers in
src/problems/and the matching documentation in the Tests section to see how diagnostic comparisons are scripted (testOrszagTang.cpp, test index). - Performance & HPC workflows. The GPU section in the installation guide plus the Running on HPC Clusters page outline how to scale to accelerators and clusters (installation guide, HPC guide).
- Contribution process. Pair the coding standards baked into
clang-tidywith the community guidance in the Contributing guide when preparing patches (and reviewCONTRIBUTING.mdat the repo root) (clang-tidy how-to, contribution guide).
Armed with the overview above, a newcomer can pick a problem driver, follow the build instructions, and iterate confidently while leaning on the documentation site for deeper dives.
Developing a new problem generator
Quokka problem generators live under src/problems/, each in its own subdirectory with a driver source file and a small CMake fragment. The sections below walk through the essential steps to bring up a new scenario, from adding the entry point to integrating it with the build and test infrastructure.
1. Create a problem skeleton
- Pick a descriptive directory name (for example
MyProblem) beneathsrc/problems/and add anadd_subdirectory(MyProblem)entry tosrc/problems/CMakeLists.txt. This is how the top-level build discovers the new problem; seesrc/problems/CMakeLists.txt. - Inside the new directory, create a
CMakeLists.txtthat uses thequokka_add_problemhelper function fromProblemHelpers.cmake. This function automatically sets up the executable (with the correct sources and CUDA compilation if needed) and optionally registers a regression test. The simplest case is:
This creates an executable namedquokka_add_problem(JOB_NAME MyProblem)MyProblemfromtestMyProblem.cpp, sets up CUDA compilation if needed, and registers a test that runs with../inputs/MyProblem.toml. To disable the test, useADD_TEST OFF:
For problems that need dimension guards or custom input files, you can combine the helper with conditional logic:quokka_add_problem(JOB_NAME MyProblem ADD_TEST OFF)
Seeif(AMReX_SPACEDIM GREATER_EQUAL 3) quokka_add_problem(JOB_NAME MyProblem) endif()src/problems/ShockCloud/CMakeLists.txtandsrc/problems/NscbcVortex/CMakeLists.txtfor examples. The helper function’s full interface is documented insrc/problems/ProblemHelpers.cmake. For problems requiring multiple tests or more complex test configurations, you may need to manually set up the executable and tests instead of using the helper. Seesrc/problems/testSN/CMakeLists.txtfor an example of a problem that requires multiple tests. - Add a driver source file (for example
testMyProblem.cpp) that will hold the problem-specific specialisations and theproblem_main()implementation described below.
2. Define problem traits
Problem generators tag their configuration by declaring an empty type (e.g., struct MyProblem { };) and specialising the trait structures that drive Quokka’s compile-time selection of physics modules.
- Every problem must specialise
Physics_Traits<MyProblem>to advertise which subsystems (hydrodynamics, radiation, MHD, etc.) are active and which unit system is in use. The linear advection driver demonstrates a minimal specialisation that disables all optional physics; examinesrc/problems/Advection/test_advection.cpp. - Problems that rely on an equation of state should also specialise
quokka::EOS_Traits<MyProblem>to provide constants such as the adiabatic index and mean molecular weight; the hydro shock tube example illustrates the pattern insrc/problems/HydroShocktube/test_hydro_shocktube.cpp.
Once the traits are in place you can specialise the Quokka or AMR simulation hooks that actually set up and evolve your scenario.
3. Provide initial conditions and runtime hooks
At a minimum, implement setInitialConditionsOnGrid for your problem’s simulation type. This routine is invoked during setInitialConditions() and must populate the cell-centred state arrays for each patch. The advection test fills a sawtooth profile by looping over the grid supplied through the helper quokka::grid struct; refer to src/problems/Advection/test_advection.cpp.
Additional hooks are available when you need them:
- Override
setCustomBoundaryConditionsif you require non-periodic inflow/outflow values, as shown by the shock tube problem described insrc/problems/HydroShocktube/test_hydro_shocktube.cpp. - Provide a
refineGridspecialisation to tag cells for AMR refinement. The shock tube driver flags zones based on the density gradient magnitude—seesrc/problems/HydroShocktube/test_hydro_shocktube.cpp. - Implement
computeReferenceSolutionwhen you want the regression harness to compare against an analytic or tabulated solution. The advection example computes an exact profile for error checking and optional plotting insrc/problems/Advection/test_advection.cpp.
Many other virtual hooks (for particles, diagnostics, derived variables, etc.) already have defaults in QuokkaSimulation, so you only need to specialise the ones your problem truly depends on.
4. Write problem_main()
The problem_main() function is the entry point that src/main.cpp calls after AMReX initialisation. It belongs in your driver file and should construct the appropriate simulation class, configure runtime parameters, and launch the run. The advection driver shows the typical flow: build boundary conditions, instantiate the simulation, adjust stopping criteria and CFL numbers, seed the initial data, and finally call evolve()—compare src/problems/Advection/test_advection.cpp and src/main.hpp.
If your problem needs exact solutions, diagnostics, or error checks, compute them before returning a status code from problem_main() so automated tests can detect failures.
5. Build and run the problem
- Regenerate or update your build tree with CMake (for example,
cmake -S . -B build -G Ninjawith the desired options). The compiled problem executables live underbuild/src/problems/<ProblemName>/once the build completes; the installation guide covers the workflow in the build instructions. - Ask CMake for the target corresponding to your new problem (e.g.,
cmake --build build --target help) and then build it with Ninja or your chosen generator. The executable name matches theJOB_NAMEyou specified inquokka_add_problem(e.g.,MyProblem), as outlined in the specific-target build section. - Run the binary from a working directory that can see your input deck, passing the
.tomlfile path as the first argument. The regression tests invoke the advection example asAdvection ../inputs/Advection.toml(note the executable name matches theJOB_NAME), which you can mimic for manual runs. Thequokka_add_problemhelper automatically configures tests to run from thetests/directory with the input file path../inputs/${JOB_NAME}.toml.
Following the steps above should give you a fully integrated problem generator that participates in Quokka’s build system and can be exercised both manually and through CTest.
Contributing Guide
Bug fixes, questions, and contributions of new features are welcome!
- Please report bugs using GitHub Issues.
- Ask questions using GitHub Discussions.
- All code contributions should be done via GitHub Pull Requests:
-
Continuous integration (CI) testing is used to ensure that any new features or changes produce the right answer and follow all of the code style guidelines.
-
A pull request (PR) should be generated from your fork of Quokka and target the
developmentbranch. (See “Git workflow” below for details.)The most important guidelines are:
- Please ensure that your PR title and first post are descriptive, since these will be used for a squashed commit message.
- Don’t group together unrelated changes in a single PR. Instead, create separate PRs for each logically-related set of changes to the code.
-
Warning
Please note: If you choose to make contributions to the code, you hereby grant a non-exclusive, royalty-free perpetual license to install, use, modify, prepare derivative works, incorporate into other computer software, distribute, and sublicense such enhancements or derivative works thereof, in binary and source code form.
The quokka developer script
The scripts/bash/quokka script provides a convenient interface for configuring, building, and running problems during development. Install it by copying it to a directory on your PATH, e.g.:
cp scripts/bash/quokka ~/.local/bin/
Commands
quokka config [-d <preset>] [--delete] [--source <file>] [--root <path>] [-D<k>=<v> ...]
Configure the CMake build directory for the given preset.
quokka build [-d <preset>] <problem> [<problem> ...] [-j <N>] [--source <file>] [--root <path>]
Compile one or more specific problem targets using ninja.
quokka build [-d <preset>] --filter <glob> [-j <N>] [--source <file>] [--root <path>]
Compile all problem targets matching a shell glob (e.g. `Rad*`).
quokka buildrun [-d <preset>] <problem> [<problem> ...] [-j <N>] [--fpe] [--input <file>] [--source <file>] [--root <path>]
Build then run one or more specific problems.
quokka buildrun [-d <preset>] --filter <pattern> [-j <N>] [--source <file>] [--root <path>]
Build matching problems then run matching tests.
quokka run [-d <preset>] <problem> [<problem> ...] [--input <file>] [-j <N>] [--fpe] [--source <file>] [--root <path>]
Run one or more problem executables from the tests/ directory.
quokka run [-d <preset>] [--filter <regex>] [-j <N>] [--source <file>] [--root <path>]
Run all tests, or those matching a regex via ctest -R.
quokka list [--root <path>]
List all available problem directories.
quokka target [-d <preset>] [--source <file>] [--root <path>]
Show all available CMake build targets.
quokka clean [--root <path>]
Remove plotfiles, checkpoints, and output files from tests/.
Presets
| Preset | Dimensionality | Build type |
|---|---|---|
1d | 1D | Release |
3d | 3D | Release |
1d-debug | 1D | Debug |
3d-debug | 3D | Debug |
Options
| Option | Description |
|---|---|
-d <preset> | Preset to use: 1d, 3d, 1d-debug, 3d-debug (default: QUOKKA_PRESET if set, otherwise 1d) |
--root <path> | Path to the quokka repository root (default: current directory) |
--input <file> | Input file for the executable (default: inputs/<problem>.toml); valid only when running exactly one <problem> |
--fpe | Enable floating-point exception traps (invalid, overflow, div-by-0) for direct executable runs (run <problem>, buildrun <problem>) |
--filter <pattern> | For run: ctest regex via ctest -R; for build: shell glob over problem names; for buildrun: build via glob and run only the tests collected from the same matched problems; exclusive with positional <problem> args. Quote patterns like 'Rad*' to avoid shell expansion before quokka sees them |
--source <file> | Optional environment file to source before config, build, buildrun, run, and target |
--delete | config only: force reconfigure by deleting existing build/<preset> directory first |
-D<k>=<v> | config only: pass extra CMake cache definitions (repeatable) |
-j <N> | Number of parallel jobs for ninja or ctest (default: 8) |
You can set QUOKKA_PRESET in your shell environment to change the default preset used by config, build, buildrun, run, and target. Passing -d <preset> still takes precedence.
build, run, and buildrun print final summary lines (<name> SUCCESS|FAIL|SKIPPED) to make tail-based status checks easy.
Typical workflow
# 1. Configure (only needed once, or after CMakeLists changes)
quokka config -d 3d
# 2. Build specific problems
quokka build -d 3d ParticleSink
quokka build -d 3d RadDust RadDustMG
# 3. Or build all matching problems
quokka build -d 3d --filter "Rad*"
# 4. Build and run in one command
quokka buildrun -d 3d RadDust RadTube
# 5. Run one problem
quokka run -d 3d ParticleSink
# 6. Run multiple problems
quokka run -d 3d RadDust RadDustMG
# 7. Run with a custom input file and FPE traps enabled
quokka run -d 3d ParticleSink --input inputs/ParticleSink_custom.toml --fpe
# 8. Run all 3D tests (8 parallel jobs)
quokka run -d 3d -j 8
# 9. Run tests matching a regex pattern
quokka run -d 3d --filter "Particle*"
# 10. List available problems (all presets)
quokka list
# 11. Clean up test output
quokka clean
Git workflow
Quokka uses git for version control. If you are new to git, you can follow one of these tutorials:
Make your own fork and create a branch on it
The basic workflow is:
- Fork the main repo (or update it if you already created it).
- Implement your changes and push them on a new branch
<branch_name>on your fork. - Create a Pull Request from branch
<branch_name>on your fork to branchdevelopmenton the main Quokka repository.
First, let us setup your local git repo. To make your own fork of the main
(upstream) repository, press the fork button on the Quokka Github page.
Then, clone your fork on your local computer. If you plan on doing a lot of Quokka development, we recommend configuring your clone to use ssh access so you won’t have to enter your Github password every time, which you can do using these commands:
git clone --branch development git@github.com:<myGithubUsername>/quokka.git
# Then, navigate into your repo, add a new remote for the main Quokka repo, and fetch it:
cd Quokka
git remote add upstream git@github.com:quokka-astro/quokka.git
git remote set-url --push upstream git@github.com:<myGithubUsername>/quokka.git
git fetch upstream
# We recommend setting your development branch to track the upstream one instead of your fork:
git branch -u upstream/development
For instructions on setting up SSH access to your Github account on a new machine, see here.
Now you are free to play with your fork (for additional information, you can visit the Github fork help page).
Note
You do not have to re-do the setup above every time. Instead, in the future, you need to update the
developmentbranch on your fork withgit checkout development git pull
Make sure you are on the development branch with
git checkout development
in the Quokka directory.
Create a branch <branch_name> (the branch name should reflect the piece
of code you want to add, like high_order_interpolation) with
git checkout -b <branch_name>
and do the coding you want. Add the files you work on to the git staging area with
git add <file_I_created> <and_file_I_modified>
Commit & push your changes
Periodically commit your changes with
git commit -m "This is a 50-char description to explain my work"
The commit message (between quotation marks) is super important in order to follow the developments and identify bugs.
For the moment, commits are on your local repo only. You can push them to your fork with
git push -u origin <branch_name>
If you want to synchronize your branch with the development branch (this is useful
when development is being modified while you are working on
<branch_name>), you can use
git pull upstream development
and fix any conflicts that may occur.
Do not merge your branch for PR into your local development branch,
because it will make your local development branch diverge from the
matching branch in the main repository after your PR is merged.
Submit a Pull Request
A Pull Request is the way to efficiently visualize the changes you made
and to propose your new feature/improvement/fix to the Quokka project.
Right after you push changes, a banner should appear on the Github page of
your fork, with your <branch_name>.
- Click on the
compare & pull requestbutton to prepare your PR. - It is time to communicate your changes: write a title and a description for
your PR. People who review your PR are happy to know
- what feature/fix you propose, and why
- how you made it (created a new class than inherits from…)
- and anything relevant to your PR (performance tests, images, etc.)
- Press
Create pull request. Now you can navigate through your PR, which highlights the changes you made.
Note
Please do not write large Pull Requests, as they are very difficult and time-consuming to review. As much as possible, split them into small, targeted PRs. For example, if you find typos in the documentation open a pull request that only fixes typos. If you want to fix a bug, make a small pull request that only fixes a bug.
If you want to implement a feature and are not sure how to split it, just open a Discussion or Issue about your plans and request feedback from other Quokka developers.
Even before your work is ready to merge, it can be convenient to create a PR (so you can use Github tools to visualize your changes). In this case, please make a “draft” PR using the drop-down menu next to the “Create pull request” button.
Once your pull request is made, we will review and potentially merge it. We recommend always creating a new branch for each pull request, as per the above instructions. Once your pull request is merged, you can delete your local PR branch with
git branch -D <branch_name>
and you can delete the remote one on your fork with
git push origin --delete <branch_name>
Generally speaking, you want to follow the following rules:
-
Do not merge your branch for PR into your local
developmentbranch that tracks Quokkadevelopmentbranch. Otherwise your localdevelopmentbranch will diverge from Quokkadevelopmentbranch. -
Do not commit in your
developmentbranch that tracks Quokkadevelopmentbranch. -
Always create a new branch based off
developmentbranch for each pull request, unless you are going to use git to fix it later.
If you have accidentally committed in development branch, you can fix it as follows,
git checkout -b new_branch
git checkout development
git reset HEAD~2 # Here 2 is the number of commits you have accidentally committed in development
git checkout .
After this, the local development should be in sync with Quokka development and your recent
commits have been saved in new_branch branch.
If for some reason your PR branch has diverged from Quokka, you can try to fix it as follows. Before you try it, you should back up your code in case things might go wrong.
git fetch upstream # assuming upstream is the remote name for the official Quokka repo
git checkout -b xxx upstream/development # replace xxx with whatever name you like
git branch -D development
git checkout -b development upstream/development
git checkout xxx
git merge yyy # here yyy is your PR branch with unclean history
git rebase -i upstream/development
You will see something like below in your editor,
pick 7451d9d commit message a
pick c4c2459 commit message b
pick 6fj3g90 commit message c
This now requires a bit of knowledge on what those commits are, which commits have been merged,
which commits are actually new. However, you should only see your only commits. So it should be
easy to figure out which commits have already been merged. Assuming the first two commits have been
merged, you can drop them by replace pick with drop:
drop 7451d9d commit message a
drop c4c2459 commit message b
pick 6fj3g90 commit message c
After saving and then exiting the editor, git log should show a clean history based on top of
development branch. You can also do git diff yyy..xxx to make sure nothing new was dropped. If
all goes well, you can submit a PR using xxx branch.
Don’t worry, if something goes wrong during the rebase, you an always git rebase --abort and start over.
Code Style Guide
Code Guidelines
Quokka developers should adhere to the following coding guidelines:
- Use curly braces for single statement blocks. For example:
for (int n=0; n<10; ++n) {
Print() << "Like this!";
}
or
for (int n=0; n<10; ++n) { Print() << "Like this!"; }
but not
for (int n=0; n<10; ++n) Print() << "Not like this.";
or
for (int n=0; n<10; ++n)
Print() << "Not like this.";
- Member variables should be suffixed with
_. For example:
amrex::Real variable_;
These guidelines should be adhered to in new contributions to Quokka, but please refrain from making stylistic changes to unrelated sections of code in your PRs.
API Documentation Using Doxygen
The Doxygen documentation is designed for advanced user-developers. It aims to maximize the efficiency of a search-and-find style of locating information. Doxygen style comment blocks should proceed the namespace, class, function, etc. to be documented where appropriate. For example:
/**
* \brief A one line description.
*
* \param[in] variable A short description of the variable.
* \param[inout] data The value of data is read and changed.
*
* A longer description can be included here.
*/
void MyFunction (int variable, MultiFab& data){
...
Additional information regarding Doxygen comment formatting can be found in the Doxygen Manual.
Flowchart
Hydro Integrator
The hydro integrator advances the conservative gas (and optionally MHD) variables that live in state_new_cc_ and the face-centered magnetic fields in state_new_fc_. The entry point is QuokkaSimulation<problem_t>::advanceHydroAtLevelWithRetries, which is called once per AMR level from AdvanceTimeStepOnLevel. The routine is designed to deliver a second-order accurate, Strang-split update while remaining robust in the presence of stiff source terms and strong shocks and rarefactions.
High-level workflow
- Source splitting: Each accepted substep begins and ends with
addStrangSplitSourcesWithBuiltin, applying the problem-specific source terms for half a timestep. - Runge–Kutta stages:
advanceHydroAtLevelimplements an SSP-RK2 integrator (with an optional first-order Euler fallback) over two stages. Stage 1 advances from the old state to an intermediate state; Stage 2 starts from the intermediate state and computes the second stage fluxes. The final update is done from the initial state using the average from the Stage 1 and Stage 2 fluxes. - Flux register coupling: When refluxing is enabled the combined RK fluxes are accumulated via
incrementFluxRegisters, guaranteeing flux conservation across AMR coarse/fine interfaces. - Tracer and dual-energy support: At each stage, face-centered velocities are used to update the auxiliary internal energy and perform a dual energy sync. Face-centered velocities produced during the update are averaged over the timestep and reused to advance tracer particles at the end of the step.
- CFL validation: A final
isCflViolatedcheck compares the accepted timestep against the maximum signal speed of the provisional updated state. Any violation forces the retry logic to reduce the timestep and redo the hydro update, ensuring the user-specifiedcflNumber_is respected.
Reconstruction and flux evaluation
advanceHydroAtLevel delegates the spatial discretisation to computeHydroFluxes:
- Primitive variables are reconstructed from the conservative state stored in the cell-centered (
state_old_cc_tmp) and, when MHD is enabled, face-centered (state_old_fc_tmp) data structures. - Multi-dimensional flattening coefficients (
ComputeFlatteningCoefficients) limit the slopes near shocks before higher-order reconstruction is applied. - Directional flux functions call the appropriate Riemann solver:
HLLCfor pure hydro,HLLDplus constrained transport (SolveInductionEqn) for MHD. The solver also returns face-centered velocities and the fastest MHD wave speeds used by the EMF update. - Before the second-order update, a first-order (donor-cell) set of fluxes is computed through
computeFOHydroFluxes. These rely on the diffusiveLLFsolvers so that first-order flux correction has the most stable possible fallback without discarding the high-order solution everywhere.
Ghost-cell requirements
The hydro update uses ghosted Riemann outputs with width nghost_Riemann. The
minimum safe value depends on two independent constraints:
- Tracer particles require
nghost_Riemann >= 2because the time-averaged face velocity used byAdvectWithUmacis stored with two ghost faces. - MHD may require additional Riemann ghosts for the EMF reconstruction and EMF averaging stencils.
For the persistent state, the current code uses:
- All hydro and MHD paths:
nghost_cc_ = nghost_Riemann + 4 nghost_fc_ = nghost_cc_
The full case table is:
| Physics path | Tracers | EMF compute | EMF averaging | nghost_Riemann | nghost_cc_ | nghost_fc_ |
|---|---|---|---|---|---|---|
| Hydro-only | off | n/a | n/a | 0 | 4 | 4 |
| Hydro-only | on | n/a | n/a | 2 | 6 | 6 |
| MHD | off | FelkerStone2017 | LondrilloDelZanna2004 | 1 | 5 | 5 |
| MHD | on | FelkerStone2017 | LondrilloDelZanna2004 | 2 | 6 | 6 |
| MHD | off | FelkerStone2017 | Balsara2025 | 2 | 6 | 6 |
| MHD | on | FelkerStone2017 | Balsara2025 | 2 | 6 | 6 |
| MHD | off | Balsara2025 | LondrilloDelZanna2004 | 1 | 5 | 5 |
| MHD | on | Balsara2025 | LondrilloDelZanna2004 | 2 | 6 | 6 |
| MHD | off | Balsara2025 | Balsara2025 | 2 | 6 | 6 |
| MHD | on | Balsara2025 | Balsara2025 | 2 | 6 | 6 |
| MHD | off | Quokka2026 | any | 3 | 7 | 7 |
| MHD | on | Quokka2026 | any | 3 | 7 | 7 |
First-order fallback and stability guards
Throughout each timestep the solver decorates the update with physics-aware stability checks:
HydroSystem<problem_t>::AddInternalEnergyPdVandPredictStepcompute the stage RHS and provisional states.redoFlagflips only when the provisional density in a cell becomes non-positive, so flagged cells have their fluxes replaced with the pre-computed first-order counterparts viareplaceFluxes/replaceEMFs. The full timestep in those cells is therefore carried out at first order in both space and time.- If any flagged cells remain after the first-order correction, the retry machinery escalates (unless
abortOnFofcFailure_is set, in which case the attempt aborts immediately). - Post-update limiters (
EnforceLimits) and optional dual-energy synchronization ensure the final conservative state obeys positivity constraints prior to CFL evaluation and refluxing.
Source terms, radiation, and coupling hooks
advanceHydroAtLevel only updates the conservative hydro variables. Radiation subcycling (subcycleRadiationAtLevel), additional operator-split source modules, and diagnostic output are invoked in the surrounding time-stepping loop.
Hydro retries algorithm
The retry loop surrounding the integrator helps prevent crashes in the presence of strong nonlinearities in the solution:
- State checkpointing: Before each attempt the routine snapshots the last accepted solution into
accepted_state_cc(andaccepted_state_fcfor MHD). Failed attempts always callrestoreHydroState()so that no partial update contaminates subsequent retries. - Adaptive substepping: Each retry chooses
total_substeps = 2^{cur_retry_level}(withcur_retry_level ≤ max_retries = 6). Progress is recorded as an integer number of base units, where one unit representsdt_lev / (2^{max_retries}). This guarantees that any retry can reinterpret previously accepted work exactly on its canonical grid. - Substep execution: For the current retry level the code computes
units_per_substep = 2^{max_retries - cur_retry_level}and evaluatesstart_substep = completed_units / units_per_substep. Only the remaining substeps are executed. The floating-point time passed toadvanceHydroAtLevelis reconstructed on demand ascurrent_substep_time = time + substep_index * dt_substep. - Partial progress handling: Accepting a substep immediately increments
completed_unitsand callsupdateAcceptedHydroState(). If a failure occurs after some substeps succeed, the retry level is bumped (capped bymax_retries) so the next attempt uses smaller substeps; a fully successful pass exits the outer loop immediately. - Failure diagnostics: Exceeding the retry budget triggers a fatal diagnostic: the code writes a
debug_hydro_state_fatalplotfile and aborts, ensuring that difficult-to-integrate states can be debugged.
Pseudocode outline
time = currentLevelTime
max_total_substeps = 1 << max_retries
accepted_state_cc = snapshotCellCenteredState()
accepted_state_fc = snapshotFaceCenteredState()
completed_units = 0
cur_retry_level = 0
while completed_units < max_total_substeps and cur_retry_level <= max_retries:
total_substeps = 1 << cur_retry_level
units_per_substep = max_total_substeps // total_substeps
start_substep = completed_units // units_per_substep
dt_substep = dt_lev / total_substeps
for substep_index in range(start_substep, total_substeps):
current_substep_time = time + substep_index * dt_substep
substep_success = advanceHydroAtLevel(current_substep_time, dt_substep)
if not substep_success:
restoreHydroState()
cur_retry_level += 1
break
completed_units += units_per_substep
updateAcceptedHydroState()
if completed_units < max_total_substeps:
writeFatalDiagnostics()
abortHydroStep()
Radiation Integrator
The radiation integrator advances the coupled radiation–matter system using the IMEX PD-ARS scheme. It is called once per AMR level from AdvanceTimeStepOnLevel, after the hydro advance, via QuokkaSimulation<problem_t>::subcycleRadiationAtLevel. Because the radiation timestep is typically much smaller than the hydro timestep (the reduced speed of light c_hat can still exceed the hydrodynamic wave speed), the radiation step is subdivided into multiple substeps within a single hydro timestep.
High-level workflow
- Substep count:
computeNumberOfRadiationSubstepsdetermines the integer number of radiation substeps needed to cover the hydro timestep at the radiation CFL number. When hydro is disabled or a constant timestep is in use, a single substep is taken. - State management: At the start of each substep after the first,
swapRadiationStatecopies the radiation hyperbolic variables fromstate_new_cc_back intostate_old_cc_, so that the integrator always has a clean “old” radiation state to advance from while the hydro variables remain instate_new_cc_. - IMEX stages per substep: Each substep applies the 3-stage IMEX PD-ARS scheme — one explicit Forward Euler stage and one explicit RK2 corrector stage, each followed by an implicit Newton–Raphson solve for the stiff matter–radiation coupling.
- Particle source injection: In 3D, stellar particles deposit their luminosity into
radEnergySourcebefore each implicit solve, giving a cell-centred luminosity density (erg s⁻¹ cm⁻³). - Flux register coupling: Radiation fluxes are accumulated into
FluxRegisters for later refluxing across AMR coarse/fine interfaces.
IMEX PD-ARS scheme
The coupled system has the form
∂U/∂t = s(U) + g(U)
where s is the stiff-explicit part (radiation transport: flux divergence) and g is the stiff-implicit part (matter–radiation exchange: emission, absorption, momentum coupling). The IMEX PD-ARS scheme integrates this with a 3-stage, 2nd-order accurate, L-stable method.
Butcher tableaux
The explicit (Aex) and implicit (Aim) tableaux are:
Explicit Aex: Implicit Aim:
c | A c | A
--+----- --+-----
0 | 0 0 0 0 | 0 0 0
1 | 1 0 0 1 | 0 1 0
1 | 1/2 1/2 0 1 | 0 1/2 1/2
|----------- |-----------
| 1/2 1/2 0 | 0 1/2 1/2
The scheme is stiffly accurate: the quadrature weights b equal the last row of A in both tableaux, so the solution at the end of each step equals the last stage value. In code the entries are defined as static constexpr members of QuokkaSimulation:
| Constant | Value | Meaning |
|---|---|---|
IMEX_Aex_21 | 1.0 | Explicit flux weight, stage 2 ← stage 1 |
IMEX_Aex_31 | 0.5 | Explicit flux weight, stage 3 ← stage 1 |
IMEX_Aex_32 | 0.5 | Explicit flux weight, stage 3 ← stage 2 |
IMEX_Aim_22 | 1.0 | Implicit solve timestep fraction, stage 2 |
IMEX_Aim_32 | 0.5 | Off-diagonal implicit weight, stage 3 ← stage 2 |
IMEX_Aim_33 | 0.5 | Implicit solve timestep fraction, stage 3 |
IMEX_alpha | 0.5 | Derived: Aim_32 / Aim_22 (Shu-Osher weight) |
Stage equations
Let U^n denote the state at the beginning of a radiation substep. Stage 1 is trivial (U^(1) = U^n). The two active stages are:
Stage 2 — predictor:
U^(2) = U^n + dt * Aex_21 * s(U^(1)) + dt * Aim_22 * g(U^(2))
The explicit part is a Forward Euler step with coefficient Aex_21 = 1; the implicit part is a backward Euler solve with dt_implicit = Aim_22 * dt = dt.
Stage 3 — corrector:
U^(3) = U^n + dt * [Aex_31 * s(U^(1)) + Aex_32 * s(U^(2))]
+ dt * [Aim_32 * g(U^(2)) + Aim_33 * g(U^(3))]
The explicit part is the standard SSP-RK2 combination of stage-1 and stage-2 fluxes; the implicit part involves the off-diagonal contribution Aim_32 * g(U^(2)) carried from stage 2, plus a new diagonal implicit solve with dt_implicit = Aim_33 * dt = dt/2.
Shu-Osher form for stage 3
The off-diagonal implicit contribution Aim_32 * g(U^(2)) does not need to be stored separately. Using the stage-2 equation to eliminate dt * g(U^(2)):
dt * Aim_22 * g(U^(2)) = U^(2) - U^n - dt * Aex_21 * s(U^(1))
Substituting into the stage-3 equation and collecting terms gives the Shu-Osher form:
let alpha = Aim_32 / Aim_22 = 0.5
U^(3)* = (1 - alpha) * U^n + alpha * U^(2)
+ dt * (Aex_31 - alpha * Aex_21) * s(U^(1))
+ dt * Aex_32 * s(U^(2))
For PD-ARS the coefficient Aex_31 - alpha * Aex_21 = 0.5 - 0.5 * 1 = 0, so no stage-1 flux divergence term appears:
U^(3)* = 0.5 * U^n + 0.5 * U^(2) + 0.5 * dt * s(U^(2))
This is exactly the SSP-RK2 corrector formula, extended to also carry forward the implicit contribution from stage 2 through the alpha * U^(2) weight.
The rest of stage 3 is a single backward Euler step: U^(3) = U^(3)* + dt Aim_33 * g(U^(3))
Implementation
Define state_xxx_gas and state_xxx_rad as the gas and radiation components of state_xxx_cc, respectively, where xxx can be new or tmp1.
Stage 1
Trivial state_new_cc_ = state_new_cc_, skipped.
Stage 2
// 1. Copy hydro-updated state into temporary (preserves gas variables)
state_tmp1_cc = state_new_cc_[lev]
// 2. Forward Euler overwrites radiation vars in state_tmp1_cc from state_old_cc_
advanceRadiationForwardEuler(..., state_tmp1_cc)
// → state_tmp1_rad = state_old_rad + dt * Aex_21 * s(state_old_rad)
// state_tmp1_gas = gas_n (unchanged by PredictStep)
// 3. Implicit solve: full Newton-Raphson with dt_implicit = Aim_22 * dt
AddSourceTerms(state_tmp1_cc, dt_implicit = Aim_22 * dt, gas_update_factor = 1.0)
// → state_tmp1 = U^(2) (radiation + gas fully updated)
Stage 3
// 4. Explicit corrector (radiation vars only — Shu-Osher form)
advanceRadiationMidpointRK2(..., state_inter = state_tmp1_cc)
// calls AddFluxesRK2(alpha=0.5, Aex_s1_coeff=0, Aex_s2_coeff=0.5)
// → state_new_rad = (1 - alpha) * state_new_rad + alpha * state_tmp1_rad
// + dt * (Aex_31 - alpha * Aex_21) * s(U^(1))
// + dt * Aex_32 * s(state_tmp1_rad)
// = 0.5 * state_new_rad + 0.5 * state_tmp1_rad + dt * 0.5 * s(state_tmp1_rad)
// 5. Shu-Osher combination for gas variables
// AddFluxesRK2 only touches radiation hyperbolic indices; gas must be combined
// explicitly via a ParallelFor kernel on components [0, nstartHyperbolic_)
// and [nstartHyperbolic_ + ncompHyperbolic_, nComp):
// state_new_gas = (1 - alpha) * state_new_gas + alpha * state_tmp1_gas
// = 0.5 * gas_n + 0.5 * state_tmp1_gas
// 6. Implicit solve of `U^(3) = U^(3)* + dt Aim_33 * g(U^(3))`: Newton-Raphson with dt_implicit = Aim_33 * dt = 0.5 * dt
AddSourceTerms(state_new_cc_, dt_implicit = Aim_33 * dt, gas_update_factor = 1.0)
// → state_new = U^(3)
Key functions
| Function | Role |
|---|---|
subcycleRadiationAtLevel | Outer loop: substep count, state swap, per-substep IMEX stages |
advanceRadiationForwardEuler | Stage 2 explicit: PredictStep with dt * Aex_21 into state_out |
advanceRadiationMidpointRK2 | Stage 3 explicit: AddFluxesRK2 with Shu-Osher coefficients, reading state_inter |
RadSystem::AddFluxesRK2 | GPU kernel: (1-alpha)*U0 + alpha*U1 + Aex_s1_coeff*F0 + Aex_s2_coeff*F1 for radiation |
RadSystem::AddSourceTermsSingleGroup | GPU kernel: Newton-Raphson implicit solve for single-group coupling |
RadSystem::AddSourceTermsMultiGroup | GPU kernel: Newton-Raphson implicit solve for multi-group coupling |
swapRadiationState | Copies radiation hyperbolic vars from state_new → state_old for next substep |
Source term interface
Both AddSourceTermsSingleGroup and AddSourceTermsMultiGroup accept explicit (dt_implicit, gas_update_factor) parameters rather than a stage integer. The caller computes:
dt_implicit = Aim_ii * dt_radiation— the effective implicit timestep for the diagonal solvegas_update_factor = 1.0— full update to all variables (no partial-update approximation)
This makes the mapping from Butcher tableau entries to solver calls transparent.
Temporary state storage
state_tmp1_cc is allocated once per call to subcycleRadiationAtLevel (before the substep loop) and reused across substeps. It holds the complete U^(2) state after stage 2, enabling the Shu-Osher combination for gas variables in step 5 above without needing to store g(U^(2)) separately.
Equivalence with the previous implementation (single-group)
Before this refactor the code used a gas_update_factor = IMEX_a32 = 0.5 trick: stage 2 applied only half the gas update, avoiding the need to store U^(2). The new implementation applies the full gas update at stage 2, then applies the Shu-Osher combination 0.5*gas_n + 0.5*gas_stage2 before stage 3’s implicit solve. The starting point for the stage 3 Newton-Raphson is identical in both cases, so for single-group radiation the numerical results are algebraically equivalent. For multi-group radiation the old gas_update_factor also entered the work-term iteration inside UpdateFlux; the new implementation with gas_update_factor = 1.0 is the mathematically correct IMEX formulation.
State Variable Component Indices
Quokka stores all cell-centred (cc) conserved quantities in a single MultiFab and all face-centred (fc) quantities in a separate per-dimension MultiFab array. The component layout within these arrays is controlled by Physics_Traits, Physics_NumVars, and Physics_Indices.
Configuration structs
Physics_NumVars
Defined in src/physics_numVars.hpp. Holds hard-coded variable counts per module.
| Constant | Value | Meaning |
|---|---|---|
numHydroVars | 6 | Conserved hydro variables: density, 3 momenta, total energy, internal energy |
numRadVarsPerGroup | 4 | Radiation variables per photon group: energy density, 3 flux components |
numDustVarsPerGroup | 4 | Dust variables per dust group: density, 3 momenta |
numMHDVars_per_dim | 1 | Face-centred MHD variables per spatial dimension (the magnetic field component normal to that face) |
numMHDVars_tot | AMREX_SPACEDIM * numMHDVars_per_dim | Total face-centred MHD variables across all dimensions |
Physics_Traits<problem_t>
Defined in src/physics_info.hpp. User-specialized per problem to enable/disable physics modules and set group counts.
| Field | Meaning |
|---|---|
is_hydro_enabled | Enable hydrodynamics |
is_radiation_enabled | Enable radiation transport |
is_dust_enabled | Enable dust dynamics |
is_mhd_enabled | Enable MHD (face-centred magnetic fields) |
is_self_gravity_enabled | Enable self-gravity |
numMassScalars | Number of mass scalars (advected proportional to density) |
numPassiveScalars | Total number of passive scalars (must be >= numMassScalars) |
nGroups | Number of radiation photon groups |
nDustGroups | Number of dust groups |
Physics module compatibility matrix
The table below summarizes the current compatibility status for the built-in physics modules documented in Quokka.
✅means the combination is supported and exercised by at least one in-tree problem or test case.❌means the combination is explicitly unsupported.⚠️means the combination appears intended to work, but no in-tree test problem currently exercises it.- A blank cell means the compatibility is currently unknown.
Here, Dust refers to the dedicated dust dynamics module enabled with Physics_Traits<problem_t>::is_dust_enabled. It is distinct from the thermal dust coupling used in several radiation test problems.
This matrix covers hydro, MHD, radiation, cooling, chemistry, particles, and the dedicated dust module. It does not include self-gravity.
| Module | Hydro | MHD | Radiation | Cooling | Chemistry | Particles | Dust |
|---|---|---|---|---|---|---|---|
| Hydro | - | ✅ | ✅ | ✅ | ✅ | ✅ | ✅ |
| MHD | - | - | ❌ | ✅ | ⚠️ | ✅ | ⚠️ |
| Radiation | - | - | - | ⚠️ | ✅ | ||
| Cooling | - | - | - | - | ⚠️ | ✅ | |
| Chemistry | - | - | - | - | - | ||
| Particles | - | - | - | - | - | - | |
| Dust | - | - | - | - | - | - | - |
Notes:
Hydro + MHDis tested by the MHD problem suite, includingAlfvenWave*,BrioWuShockTube,MHDBlast,MHDQuirk, andOrszagTang.Hydro + radiationis tested by the radiation-hydrodynamics problems such asRadhydroPulse*,RadhydroShock*,RadhydroShell,RadTube, andRadhydroBB.Hydro + coolingis tested byResampledCoolingTest,ShockCloud,RandomBlast, andSN.Hydro + chemistryis tested byPrimordialChemandPopIII.MHD + radiationis explicitly disabled insrc/QuokkaSimulation.hppwithstatic_assert(!(is_mhd_enabled && is_radiation_enabled), "MHD + Radiation is not supported yet.").MHD + coolingis exercised by theSNproblem withis_mhd_enabled = trueandcooling.enabled = 1.Hydro + particlesis exercised by problems such asBinaryOrbitCIC,ParticleSink*,ParticleSF,ParticleRadiation,RandomBlast, andSN.MHD + particlesis exercised byDiskGalaxy,ParticleAccretion,ParticleCreation,ParticleSink, andParticleSinkFormation.Radiation + particlesis exercised byParticleRadiationandGravRadParticle3D.Cooling + particlesis exercised byParticleSF,TallBoxSf,RandomBlast, andDiskGalaxythrough their input files.Hydro + dustis exercised by the dust dynamics problemsDustAdvection*,DustDamping*,DustSoundwave, andDustyShock.MHD + dustis marked untested: the drag implementation contains MHD-aware code paths, but there is no in-tree problem that enables bothis_mhd_enabledandis_dust_enabled.
Cell-centred state vector layout
All cell-centred conserved variables are packed into a single MultiFab (state_new_cc_). The total number of components is Physics_Indices<problem_t>::nvarTotal_cc.
Physics_Indices<problem_t>
Defined in src/physics_info.hpp. Computes starting indices and total component counts.
| Constant | Definition | Meaning |
|---|---|---|
nvarTotal_cc_adv | 1 | Number of cc variables for a pure advection problem (no hydro, no radiation) |
nvarTotal_cc | (see below) | Total number of cell-centred components in the state vector |
hydroFirstIndex | 0 | Starting index of hydro variables |
pscalarFirstIndex | numHydroVars (= 6) | Starting index of passive scalars |
dustFirstIndex | pscalarFirstIndex + numPassiveScalars | Starting index of dust variables |
radFirstIndex | dustFirstIndex + numDustVarsPerGroup * nDustGroups * is_dust_enabled | Starting index of radiation variables |
nvarPerDim_fc | numMHDVars_per_dim * is_mhd_enabled | Number of face-centred variables per dimension |
nvarTotal_fc | AMREX_SPACEDIM * nvarPerDim_fc | Total face-centred variables |
mhdFirstIndex | 0 | Starting index of MHD variables within each face-centred MultiFab |
nvarTotal_cc calculation:
- If neither hydro nor radiation is enabled:
nvarTotal_cc = nvarTotal_cc_adv(= 1, for pure advection). - Otherwise:
nvarTotal_cc = numHydroVars + numPassiveScalars + numDustVarsPerGroup * nDustGroups * is_dust_enabled + numRadVarsPerGroup * nGroups * is_radiation_enabled.
Cell-centred component map
The cell-centred state vector is laid out contiguously as:
Index: 0 6 6+Nps dustFirstIndex radFirstIndex
|--- Hydro (6) ------------|-- Scalars -----|--- Dust ------|- Radiation ---------|
Hydro block (6 variables, starting at hydroFirstIndex = 0)
Defined by HydroSystem<problem_t>::consVarIndex:
| Index | Name | Quantity |
|---|---|---|
| 0 | density_index | Gas mass density |
| 1 | x1Momentum_index | x-momentum |
| 2 | x2Momentum_index | y-momentum |
| 3 | x3Momentum_index | z-momentum |
| 4 | energy_index | Total energy density |
| 5 | internalEnergy_index | Auxiliary internal energy (dual energy formalism) |
Passive scalar block (numPassiveScalars variables, starting at pscalarFirstIndex = 6)
| Index | Name | Quantity |
|---|---|---|
| 6 .. 6+Nms-1 | scalar0_index | Mass scalars (partial densities; sum must equal total density; renormalized if not) |
| 6+Nms .. 6+Nps-1 | Ordinary passive scalars (arbitrary quantities; no constraints) |
Explanation:
Here, Nms = numMassScalars and Nps = numPassiveScalars.
- The first
numMassScalarsentries in the passive scalar block correspond to mass scalars. Mass scalars represent partial densities whose sum should equal the total density at each cell, and they will be renormalized if this condition is not met. - Any remaining entries (
Nps - Nms) are ordinary passive scalars, which store arbitrary advected quantities and are not required to satisfy any normalization constraint. - If
numMassScalars = 0, then all components in the passive scalar block are ordinary passive scalars.
Dust block (numDustVarsPerGroup * nDustGroups variables, starting at dustFirstIndex)
Only present when is_dust_enabled = true. For each dust group g (0-indexed):
| Offset within group | Name | Quantity |
|---|---|---|
| 0 | dustDensity_index | Dust density |
| 1 | x1DustMomentum_index | Dust x-momentum |
| 2 | x2DustMomentum_index | Dust y-momentum |
| 3 | x3DustMomentum_index | Dust z-momentum |
Variables for group g start at dustFirstIndex + g * numDustVarsPerGroup.
Radiation block (numRadVarsPerGroup * nGroups variables, starting at radFirstIndex)
Only present when is_radiation_enabled = true. For each photon group g (0-indexed):
| Offset within group | Name | Quantity |
|---|---|---|
| 0 | radEnergy_index | Radiation energy density |
| 1 | x1RadFlux_index | Radiation flux, x-component |
| 2 | x2RadFlux_index | Radiation flux, y-component |
| 3 | x3RadFlux_index | Radiation flux, z-component |
Variables for group g start at radFirstIndex + g * numRadVarsPerGroup.
Note: RadSystem defines nstartHyperbolic_ = radFirstIndex and nvarHyperbolic_ = numRadVarsPerGroup * nGroups. Its internal nvar_ includes all preceding variables: nvar_ = nstartHyperbolic_ + nvarHyperbolic_ (equal to nvarTotal_cc).
Face-centred state vector layout
Face-centred variables are stored in a std::array<MultiFab, AMREX_SPACEDIM> (state_new_fc_), one MultiFab per spatial dimension. Each contains nvarPerDim_fc components.
MHD block (1 variable per dimension, starting at mhdFirstIndex = 0)
Only present when is_mhd_enabled = true. Defined by MHDSystem<problem_t>::varIndex_perDim:
| Index (per dim) | Name | Quantity |
|---|---|---|
| 0 | bfield_index | Normal component of the magnetic field on that face |
The face-centred storage follows the Constrained Transport (CT) convention: state_new_fc_[0] stores Bx on x-faces, state_new_fc_[1] stores By on y-faces, and state_new_fc_[2] stores Bz on z-faces (in 3D).
Per-system variable counts
Each physics system class also defines its own nvar_ for internal use:
| Class | nvar_ | Meaning |
|---|---|---|
HydroSystem | numHydroVars + numPassiveScalars + numDustVarsPerGroup * nDustGroups * is_dust_enabled | All cc variables managed by the hydro solver (hydro + scalars + dust) |
RadSystem | radFirstIndex + numRadVarsPerGroup * nGroups | All cc variables up to and including radiation (= nvarTotal_cc) |
MHDSystem | nvar_per_dim_ = 1 per dimension, nvar_tot_ = AMREX_SPACEDIM total | Face-centred magnetic field components |
Example: typical hydro + radiation problem
For a problem with numMassScalars = 1, numPassiveScalars = 2 (1 mass scalar + 1 additional passive scalar), is_dust_enabled = false, nGroups = 2, is_radiation_enabled = true:
Cell-centred layout (16 components total):
[0] density
[1] x1Momentum
[2] x2Momentum
[3] x3Momentum
[4] energy
[5] internalEnergy
[6] scalar0 (mass scalar)
[7] scalar1 (passive scalar)
[8] radEnergy (group 0)
[9] x1RadFlux (group 0)
[10] x2RadFlux (group 0)
[11] x3RadFlux (group 0)
[12] radEnergy (group 1)
[13] x1RadFlux (group 1)
[14] x2RadFlux (group 1)
[15] x3RadFlux (group 1)
Debugging
General guidelines
A good general guide to debugging simulation codes is provided by WarpX (although some details are WarpX-specific).
Common bugs
You might be reading this page because you have encountered a common type of bug:
-
This is when the code accesses an array with an index that corresponds to an element that doesn’t actually exist. In C++, this causes the computer to access a memory location that is completely unrelated to the array that you intended to access. In a CPU code, this usually causes a silently incorrect result, but on GPU, this may actually cause the simulation to crash. However, if you are accessing an
amrex::Array4object and you have compiled in Debug mode, then AMReX will issue an error message when this occurs. There is a significant performance cost to this error checking, so it does not occur when compiled in Release mode. -
accessing a host variable from the GPU
The second most common type of bug encountered in Quokka is accessing a host variable (i.e., a variable that can only be accessed from code that runs on the CPU) from code running on the GPU (i.e., within a
ParallelFor). Sometimes the compiler will detect this situation and print an error message, but often this will only present an issue when actuallly running the code – for instance, this can happen when the GPU code tries to dereference a pointer to an address in CPU memory. In that case, the only way to debug this error is to run Quokka undercuda-gdb(or, on AMD GPUs,rocgdb).
How to debug on GPUs
The best way to debug on GPUs is to... not debug on GPUs. That is, it is always easier to instead debug the problem on a CPU-only run. GPU debugging is very painful and itself quite buggy. This is unfortunately true for all GPU vendors.
- Try to shrink the problem to a size that can run on a single node, or (even better) a single MPI rank / CPU core, but where it still exhibits the error that you are trying to debug.
- Build Quokka without GPU support but with
-DCMAKE_BUILD_TYPE=Debugand re-run. If there are any array out-of-bounds errors, it will stop and report exactly which array is being accessed out-of-bounds and what the indices are. The only downside is that Quokka will run very slowly in this mode.- For more details, see the AMReX debugging guide.
- Build Quokka without GPU support but with
-DCMAKE_BUILD_TYPE=Release -DENABLE_ASAN=ON. This turns on the AddressSanitizer, which checks for out-of-bounds array accesses and other memory bugs. This is faster than the previous method, but it produces less informative error messages (e.g., no array indices).- This method may produce a lot of messages about memory leaks, which are not necessarily bugs, and should not cause GPU crashes. These messages can be disabled if you are looking for, e.g., out-of-bounds array accesses, which is a class of bug that can cause a GPU crash.
- It is recommended to set these environment variables when you run it:
ASAN_OPTIONS=abort_on_error=1:fast_unwind_on_malloc=1:detect_leaks=0 UBSAN_OPTIONS=print_stacktrace=0. Note that CTest appends its own options to this environment variable when running tests, so it is recommended to run the simulation manually (i.e., withoutmake test,ninja test, orctest). - For more information, see this guide to using AddressSanitizer in an HPC context.
- On AMD GPUs, there is a GPU-aware AddressSanitizer. Currently, enabling this requires manually changing the compiler flags.
How to actually debug on GPUs
As an absolute last resort if it is impossible to reproduce the error you are seeing on a CPU-only run, then the best option is to:
- downsize the simulation to fit on a single GPU
- start the simulation on an NVIDIA GPU from within CUDA-GDB (see the CUDA-GDB documentation and slides).
- hope CUDA-GDB does not itself crash
- hope CUDA-GDB produces a useful error message that you can analyze
NVIDIA also provides the compute-sanitizer tool that is essentially the equivalent of AddressSanitizer (see the ComputeSanitizer documentation). Unfortunately, it does not work as reliably as AddressSanitizer, and may itself crash while attempting to debug a GPU program.
For AMD GPUs, you have to use the AMD-provided debugger rocgdb. A tutorial its use is available here.
AMD also provides a GPU-aware AddressSanitizer that can be enabled when building Quokka. Currently, the compiler flags must be manually modified in order to enable this. For details, see its documentation.
GPU kernel asynchronicity
By default, GPU kernels launch asynchronously, i.e., execution of CPU code continues before the kernel starts on the GPU. This can cause synchronization problems if there is an implicit assumption about the order of operations with respect to CPU and GPU code.
The easiest way to debug this is to set the environment variables:
CUDA_LAUNCH_BLOCKING=1on NVIDIA GPUs, orHIP_LAUNCH_BLOCKING=1on AMD GPUs.
This will cause the CPU to wait until the GPU kernel execution is complete before continuing past the call to ParallelFor.
For more details, refer to the AMReX GPU debugging guide.
Testing for GPU race conditions
GPU race conditions can cause non-deterministic behavior where the same simulation produces different results on different runs, or crashes only intermittently. To systematically test for race conditions, use the provided test script:
./scripts/bash/test_gpu_race.sh -b <binary> -i <input_file> -n <max_timesteps>
Parameters:
-b <binary>: Path to the test binary (e.g.,./build/src/FieldLoop/test_field_loop)-i <input_file>: Path to the input file (e.g.,inputs/FieldLoop.toml)-n <max_timesteps>: Maximum number of timesteps to run
Example:
./scripts/bash/test_gpu_race.sh -b ./build/src/FieldLoop/test_field_loop -i inputs/FieldLoop.toml -n 10
How it works:
- Runs the same simulation twice - once with
CUDA_LAUNCH_BLOCKING=1(synchronous) and once withCUDA_LAUNCH_BLOCKING=0(asynchronous) - Compares the final plotfiles using AMReX’s
fcomparetool with zero tolerance - If the results differ, this indicates a race condition where kernel execution order affects the outcome
Interpreting results:
- SUCCESS: Plotfiles are identical - no race condition detected
- FAILURE: Plotfiles differ or non-blocking run crashes - race condition detected
The script preserves temporary files for analysis when a race condition is found. These can be examined to understand the nature of the differences between the two runs.
When all else fails: Debugging with printf
If you have tried all of the above steps, then you have to resort to adding printf statements within the GPU code. Note that printf inside GPU code is different from the CPU-side printf function, as explained in the NVIDIA documentation.
Assertions and error checking
AMReX assert macros
AMReX provides several assertion macros:
AMREX_ASSERT: Enabled when either-DAMREX_DEBUGor-DAMREX_USE_ASSERTIONis defined. When enabled, it callsamrex::Assert()which will abort unless both-DNDEBUGis defined AND-DAMREX_USE_ASSERTIONis NOT defined. This means:- The macro itself is a no-op when neither
AMREX_DEBUGnorAMREX_USE_ASSERTIONis defined - When the macro is enabled and assertion fails, it will abort unless
NDEBUGis defined withoutAMREX_USE_ASSERTION
- The macro itself is a no-op when neither
AMREX_ALWAYS_ASSERT: Always callsamrex::Assert()regardless of build configuration. The actual abort behavior follows the same rules asAMREX_ASSERT- it will abort unless both-DNDEBUGis defined AND-DAMREX_USE_ASSERTIONis NOT defined. Note that CMake adds “-DNDEBUG” by default when “CMAKE_BUILD_TYPE=Release”. (See this GitHub discussion for details.)
Abort
Because the default CMake flags added in Release mode causes AMREX_ALWAYS_ASSERT not to function in GPU code, amrex::Abort is the best option to use if you want to abort a GPU kernel.
amrex::Abort requires additional GPU register usage, so it should be used sparingly. The best strategy for error handling is often to set a value in an array that indicates an iterative solve failed in a given cell. (This is what Castro does for its nuclear burning networks.)
For more details, see the AMReX documentation on assertions and error checking.
Performance tips
Prerequisites
You should:
- Understand what a GPU kernel is. (For reference, consult these notes.)
- Understand what a processor register is.
- Know that calling
amrex::ParallelForlaunches a GPU kernel (when GPU support is enabled at compile time).
GPU hardware characteristics
GPUs have hardware design features that make their performance characteristics significantly different from CPUs. In practice, two factors dominate GPU performance behavior:
- Kernel launch latency: this is a fundamental hardware characteristic of GPUs. It takes several microseconds (typically 3-10 microseconds, but it can vary depending on the compute kernel, the GPU hardware, the CPU hardware, and the driver) to launch a GPU kernel (i.e., to start running the code within an
amrex::ParallelForon the GPU). In practice, latency is generally longer for AMD and Intel GPUs. - Register pressure: the number of registers per thread available for use by a given kernel is limited to the size of the GPU register file divided by the number of threads. If a kernel needs more registers than are available in the register file, the compiler will “spill” registers to memory, which will then make the kernel run very slowly. Alternatively, the number of concurrent threads can be reduced, which increases the number of registers available per thread.
- For more details, see these AMD website notes and OLCF training materials.
MPI communication latency vs. bandwidth
A traditional rule of thumb for CPU-based MPI codes is that communication latency often limits performance when scaling to large number of CPU cores (or, equivalently, MPI ranks). We have found that this is not the case for Quokka when running on GPU nodes (by, e.g., adding additional dummy variables to the state arrays).
Communication performance appears to (virtually always) be bandwidth limited. There are two likely reasons for this:
- GPU node performance is about 10x faster than CPU node performance, whereas network bandwidth is only 2-4x larger on GPU nodes compared to CPU nodes. The network bandwidth to compute ratio is therefore lower on GPU nodes than on CPU nodes.
- GPU kernel launch latency (3-10 microseconds) is often larger than the minimum MPI message latency (i.e., the latency for small messages to travel between nodes) of 2-3 microseconds.
Guidelines
- Combine operations into fewer kernels in order to reduce the fraction of time lost to kernel launch latency
- This can also be done by using the
MultiFabversion ofParallelForthat operates on all of the FABs at once, rather than launching a separate kernel for each FAB. This should not increase register pressure. - However, combining multiple kernels can increase register pressure, which can decrease performance. There is no real way to know a priori whether there will be a net performance gain or loss without trying it out. The strategy that yields the best performance may be different for GPUs from different vendors!
- This can also be done by using the
- Split operations into multiple kernels in order to decrease register pressure
- However, this may increase the time lost due to kernel launch latency. This is an engineering trade-off that must be determined by performance measurements on the GPU hardware. This trade-off may be different on GPUs from different vendors!
- In order to decrease register pressure, avoid using
printf,assert, andamrex::Abortin GPU code . All of these functions require using additional registers that could instead be allocated to the useful computations does in a kernel. This may require a significant code rewrite to handle errors in a different way. (You should not just ignore errors, e.g. in an iterative solver.) - Experts only: Manually tune the number of GPU threads per block on a kernel-by-kernel basis. This can reduce register pressure by allowing each thread to use more registers. Note that this is an advanced optimization and should only be done with careful performance measurements done on multiple GPUs. The AMReX documentation provides guidance on how to do this.
How to use clang-tidy
clang-tidy is a command-line tool that automatically enforces certain aspects of code style and provides warnings for common programming mistakes. It automatically runs on every pull request in the Quokka GitHub repository.
Using clang-tidy with VSCode
The easiest way to use clang-tidy on your own computer is to install the clangd extension for Visual Studio Code (VSCode).
(VSCode itself can be downloaded here.)
Command-line tools
The clangd extension for VSCode might not check all warnings, so you can also run clang-tidy from the command line (see the documentation). Here is a minimal example of how to use it:
clang-tidy src/particles/* -p ./build
which will run clang-tidy on all the files in the src/particles directory.
To see the clang-tidy warnings that are relevant only to the code changes you’ve made, you can use scripts/tidy.sh. Example usage:
./scripts/tidy.sh ./build
This will run clang-tidy on all the files that have been modified with respect to the previous commit.
./scripts/tidy.sh ./build previous
This will run clang-tidy on all the files that have been modified in the last commit.
./scripts/tidy.sh ./build origin
This will run clang-tidy on all the files that have been modified with respect to the remote branch.
./scripts/tidy.sh ./build dev
This will run clang-tidy on all the files that have been modified with respect to the local development branch.
To see the clang-tidy warnings that are relevant only to individual lines of code you’ve changed, you can use the clang-tidy-diff.py Python script.
Maintainers Guide
Quokka maintainers safeguard long-term technical direction and scientific integrity for a multi-physics AMReX application that must build across multiple toolchains and hardware platforms. You review pull requests (PRs = proposed code changes) to ensure they are correct, efficient, portable (CPU/GPU; CUDA or HIP; x86 or ARM; macOS or Linux), and reproducible (others can repeat results).
Important: Stability expectations
- Problem-file interface may change occasionally to support new features.
- Runtime
ParmParseoptions should change rarely.- Output file format must never change.
Tip: Review tone
Be explicit, patient, and educational. Assume the author is still learning; explain what to change and why with short examples or links.
Note: Long-term stewardship
Publication deadlines or short-term milestones must never override the mandate to protect Quokka’s scientific integrity and technical direction. When in doubt, slow down, request revisions, or defer merges until the change meets the bar.
For quick definitions of terms used throughout this guide, see the Glossary.
Intake & Triage
-
Require a complete PR description (plain language).
- Must include:
- Problem: what bug/feature/refactor is addressed.
- Approach: algorithms; units (e.g., cgs); error tolerances (e.g., relative error ≤ 1e-8); boundary conditions (periodic, inflow, outflow, reflecting; Dirichlet, Neumann, or periodic for elliptic solvers); staggering (cell-centered vs. face-centered).
- Stability impact: does it touch
ParmParseoptions, the output file format, or the problem-file interface? (Explain how.) - Performance results (if applicable): before/after timing or brief profiling on a test problem.
- Reproducer steps (if applicable): exact build options (CMake),
ParmParseinputs, MPI process count, and GPU backend (CUDA or HIP).
- Must include:
-
Target the right branch and scope. PRs should come from a fork/feature branch and target
development. Looking at the last 100 squash merges and filtering out trivial updates (<50 changed lines, e.g., submodule bumps or workflow tweaks), a typical PR lands around ~238 changed lines across four files (75th percentile ≈357 lines, 90th percentile ≈1.1k). Use that baseline: if a PR mixes unrelated changes, sprawls beyond ~8–10 files, or pushes well past ~400–500 lines of real code delta, ask for smaller, topic-focused PRs (“one PR = one story”).
Correctness & Numerics
-
Tests are mandatory.
- Provide:
- Unit/algorithm test: checks a single function or kernel. Currently, there are no unit tests in Quokka.
- Physics test: a small simulation test problem that runs in a few minutes. All tests in Quokka are currently physics tests.
- CI coverage: our GitHub Actions matrix drives
canary runagainst curatedctestlists on Linux/gcc (Release + ASAN), macOS/AppleClang, and ARM64 (Debug, Release, HWASan) plus two MHD-focused suites. Wire new problems into those lists (orregression/quokka-tests.inifor longer inputs) so they execute automatically. The GitHub Actionshipjob is compile-only; runtime HIP coverage comes from/azp runon the Azure Pipelines self-hosted AMD GPUs. The Azure Pipelines CI also tests the CUDA backend on an NVIDIA GPU. Note any additional backend-specific manual validation you perform.
- Provide:
-
Document numerical expectations in code/comments. State valid input ranges, units, absolute/relative tolerances, conditioning notes (sensitivity to small input changes), staggering (cell-centered vs. face-centered), and the required number of ghost cells for correctness.
-
Boundary conditions and conservation (if applicable). Describe how boundary conditions are applied and show that conserved quantities (mass, momentum, energy) remain correct within tolerance (norms or integrals over the mesh data).
Documentation Policy
-
All PRs that add new features must include documentation in the Markdown
docs/folder. -
New physics features must be marked in the documentation as “beta” if they have not been fully tested in a published scientific application. This is intended to warn users to exercise caution if they choose to use features that are not yet thoroughly tested. When a science paper is published, a new PR should be submitted to remove this marking.
Stability Policy
-
Runtime
ParmParseoptions (change rarely).- Default stance: options are stable; avoid renaming/removing.
- If change is necessary:
- Provide automatic compatibility when feasible (accept old name, map to new, and warn).
- Emit clear deprecation warnings that name the replacement and include a removal date/version.
- Document the change prominently (release notes; “Upgrading” section).
- Use the ADR process (below) for any non-trivial change in behavior or naming.
-
Output file format (must never change).
- Do not modify existing structure, metadata, or semantics.
- If new data must be written, use additive outputs that do not alter or invalidate existing files (e.g., auxiliary files or parallel sidecar outputs) so existing analysis tools remain valid.
- Any proposal that risks incompatibility should be rejected outright or re-scoped to preserve the current format.
-
Core hydro integrator (
QuokkaSimulation::advanceHydroAtLevel).
Danger: Critical change protocol
- Treat every edit as **critical**: require an ADR with numerical justification, targeted regression coverage spanning CPU and at least one GPU backend, and explicit before/after plots or diagnostics for representative hydro problems. - Demand quantified performance data (solver wall time, per-stage timing, halo exchange cost) and call out any tolerable slowdowns or accuracy gains. - Double-check plotfile contents and conserved quantities for regressions before approving.
- Problem-file interface (may change occasionally).
- Changes are acceptable to support new physics/modules, but must be intentional and documented.
- Require an ADR describing the motivation, alternatives, and migration plan.
- Migrate all in-tree problem files in the same PR; out-of-tree problems are not supported and must supply their own migrations.
Specific Pitfalls to Avoid
Warning: Watch for these regressions-in-waiting
- Renaming widely-used variables. Mechanical symbol churn often leaves call sites or device code untouched, introducing subtle bugs in solver logic and diagnostics.
- Adding redundant physics implementations. Multiple solvers for the same physical process explode our validation matrix; prefer extending or generalising existing kernels.
- Copy/pasting shared routines. Duplicated flux or source-term code drifts out of sync; factor shared helpers instead so fixes land once.
Reproducibility
-
Define the reproducibility level and test it.
- Bitwise identical: outputs match exactly.
- Within a tolerance: values may differ slightly but within bounds.
- Statistically equivalent: stochastic runs match in distribution with fixed seeds. Control randomness with documented seeds; avoid rank-dependent draw order.
-
Record run metadata (log/output). Always include: git commit hash; AMReX version; compiler and GPU backend (CUDA or HIP) + versions; MPI version; GPU model; key build flags; full
ParmParsedump; and domain decomposition. Keep minimal input decks with tests so others can rerun them. Plotfiles and checkpoints automatically persist provenance tometadata.yaml, which recordsquokka_version,git_hash_quokka,git_hash_amrex, and the current unit and constant blocks by default. Update or extend that node (viasimulationMetadata_) when additional diagnostics are essential for reruns.
CI Gates
- Automated checks must stay green (GitHub Actions).
CMake: Ubuntu + GCC 11 Release with-DENABLE_ASAN=ON; builds the full 3D configuration and runs the maincanarysuite.CMake (macOS): macOS 14 + Apple Clang Release; runs the samecanarylist.linux-arm64-{debug,release,HWASan}: ARM runners using Clang; run fast 1D smoke tests in Debug, Release, and HWASan configurations.warnings: Ubuntu build with-DWARNINGS_AS_ERRORS=ON; fails on any new compiler warning.MHD-WaveTestsandMHD-WaveTests-ASAN: GCC Release and ASAN builds that rebuild only MHD-labelled targets and execute the MHD-focusedcanarysubsets.hip: ROCm/clang toolchain compile of the full codebase (GitHub Actions build-only sanity check).OpenPMD: builds withQUOKKA_OPENPMD=ONand runs the 3D blast problem under MPI to exercise ADIOS2/openPMD output.- Static analysis & lint:
clang-tidy-review(with follow-up comment job) andcodespell. - Docs & metadata:
docs(mdBook build),mdBook validation,dependency-review,codeql, andscorecard. - Azure Pipelines (self-hosted GPUs): manual
/azp runtrigger by a maintainer launches CUDA and HIP runtime tests on secured NVIDIA/AMD runners; required for accelerator-sensitive changes. - Nightly regression suite: runs the
regression/harness against reference plotfiles on a separate pipeline; failures mean the gold data and code outputs diverged and need investigation. - Most jobs go through
check_changes.yml; docs-only or workflow-only PRs will auto-skip compute-heavy runs, so double-check locally if you bypass CI by rebasing after review.
Tip: Docs-only changes
When automation skips builds, spot-check formatting and links locally (`./scripts/bash/docs_build.sh`) before approving to avoid post-merge surprises.
Architecture Decision Records
Use Architecture Decision Records for high-impact or hard-to-reverse changes—especially anything touching ParmParse options, the problem-file interface, or output formats. They capture competing options, supporting evidence, and the rollout plan so we can resolve debates quickly and revisit choices with context.
Merge Policy
Warning: Need to undo a merge?
See the dedicated PR Revert Policy for decision criteria and the step-by-step workflow.
- Be explicit, kind, and disciplined.
- Mark comments as blocking (must fix before merge) or non-blocking (suggestions).
- If discussion becomes subjective (naming/style), time-box and move to an ADR.
- Timelines:
- Acknowledge a new PR within 5 business days.
- Provide a first substantive review within 14 days.
- For high-risk or broadly user-facing changes (
ParmParse, output format, problem-file interface), require two maintainer approvals; otherwise one is sufficient. - Use squash-and-merge by default (keeps history clean).
Architecture Decision Records (ADRs)
Note: Purpose
ADRs document important technical decisions so future contributors understand the problem we solved, the options we evaluated, and how to revisit the choice if assumptions change.
When to Author an ADR
Write an ADR whenever a change has broad impact, is hard to reverse, or triggers sustained debate.
Warning: Common triggers
- Adjusting
ParmParseoptions beyond trivial additions (renames, removals, behaviour changes).- Any risk to the output file format (typically rejected; document how compatibility is preserved if discussed).
- Modifying the problem-file interface (new hooks, migration plans, deprecations).
- Shifts in accuracy/performance trade-offs (limiters, fluxes, numerical precision), or non-trivial dependency/toolchain updates.
ADR Paths
PR-Initiated ADR (resolving review disagreements)
- Distil blocking PR comments into 3–5 explicit questions.
- Create
docs/adrs/ADR-xxxx-title.mdand link it from the PR. - Assign 2–3 maintainers (include numerics + GPU expertise) as reviewers.
- Provide evidence: options/trade-offs, CPU + CUDA/HIP benchmarks, accuracy/tolerance notes, portability, migration impact.
- Time-box discussion to ≤10 business days; schedule an optional 30–45 minute design call if needed.
- Decide and tag the ADR: Accepted / Rejected / Superseded / Amended; reference PRs/issues.
- Apply the outcome: update the PR, CI, documentation, and deprecation/migration notes.
Proactive ADR (not tied to a PR)
- Open an issue labeled
type:ADR-proposaloutlining Problem, Goals/Non-goals, Affected modules, and Risks. - Draft the ADR in
docs/adrs/ADR-xxxx-title.md; include a short rollout roadmap (milestones, owners). - Set roles: Driver (author), Reviewers (numerics + GPU/AMReX), Consulted (downstream users if affected).
- Add prototypes or microbenchmarks when helpful (CPU + CUDA/HIP).
- Leave a ≤21 day comment window (to accommodate academic schedules) or close earlier with explicit sign-off.
- Decide and define the rollout: update CI coverage, add tests, document migrations (especially for problem interfaces or
ParmParseadjustments). - Maintain the ADR lifecycle: revisit annually or when assumptions change, preferring supersession over silent drift.
ADR Template
# ADR-XXXX: <Short Title>
Date: <YYYY-MM-DD> • Status: Proposed | Accepted | Rejected | Superseded | Amended
## Context
Problem, constraints (MPI, CUDA/HIP), affected modules and users.
Touches: ParmParse options? Output format? Problem-file interface?
## Options
- Option A: …
- Option B: …
(Include small benchmarks: CPU, CUDA, HIP; accuracy/tolerance notes.)
## Decision
Chosen option and rationale.
## Consequences
Performance/accuracy impact, portability (CUDA/HIP), maintenance cost,
migration plan (deprecations, updates for problem-file interfaces; NO output-format changes).
## Rollout & Testing
CI matrix updates (CUDA/HIP/CPU), new tests, deprecations, docs updates.
## Links
PRs, issues, design notes, prior ADRs.
Ending PR Deadlocks with ADRs
Move prolonged or subjective debates into an ADR. After a decision:
- Mark PR threads resolved or superseded by ADR.
- Merge only after the ADR is Accepted and CI/tests reflect the choice.
Pull Request Revert Policy
Warning: Purpose
Reverts protect users, safeguard maintainability, and preserve scientific integrity when a merged change causes regressions that cannot wait for a fix-forward. They are not punitive—they provide a fast, low-risk way to restore a reliable baseline so contributors can investigate and reintroduce the change safely.
When to Revert
Trigger a revert only for serious issues in parts of the code that are not marked as “beta”:
Danger: Typical revert triggers
- Major correctness bugs (e.g., broken conservation, highly unstable solver, incorrect physics equations).
- Significant performance regressions on maintained benchmarks (typically >10–15% slower without an acceptable trade-off).
- Broken builds or CI on supported targets (CPU, CUDA, HIP) that cannot be repaired quickly. This applies even for beta features.
- Incompatible output-format changes that break existing plotfile readers.
- Unreproducible outputs or race conditions exposed by deterministic reruns or sanitizer tooling.
- Security or licensing concerns (e.g., unsafe dependency, incompatible license).
Revert Workflow
- Open an issue summarising impact, affected versions, and how to reproduce. Label it
regression,revert-candidate, and addbackport-neededif release branches are affected. - Prepare the revert PR:
- Title it
Revert: #<original-PR> <original title>. - Use
git revert <merge-commit-sha>when possible to keep history clean. - Keep the change minimal—no refactors or unrelated fixes.
- Add a failing test that reproduces the problem (fails on
development, passes after the revert).
- Title it
- Explain clearly in the PR body: user impact, scope, why revert beats a quick fix-forward, and the plan to reintroduce safely if needed.
- Review & merge quickly—aim for ≤3 business days once consensus forms. One maintainer approval is enough for low-risk internal fixes; require two approvals when the original change was high risk or broadly user-facing.
After the Revert
Note: Post-merge checklist
- File a follow-up issue to fix forward: root cause, new tests, ADR if architecture changes are involved.
- Note the revert in the changelog and backport to supported release branches if users rely on the affected feature.
- For
ParmParseor problem-file interface changes, publish a migration guide before attempting re-introduction.
Guiding Principles
Tip: Guiding principles
- Prefer a temporary revert plus a clear follow-up plan over leaving the code broken.
- Always attach evidence (inputs, logs, benchmarks) to justify the revert.
- Keep the revert PR surgical and traceable to the original change.
- Treat reverts as a collaborative safety measure; focus on the fix, not assigning blame.