Postprocessing
There are several ways to post-process the output of Quokka simulations. AMReX PlotfileTools, yt, and VisIt all allow you to analyze the outputs after they are written to disk.
Amrvis-container
Amrvis-container bundles Amrvis in a Docker/Apptainer image with a browser-based X11 frontend. To browse Quokka plotfiles locally (Docker required), run from the Amrvis-container repo:
./launch_amrvis_browser.sh /path/to/plotfiles
The target directory is bind-mounted to /home/vscode/data in the container. The launcher prints a one-time password; open http://localhost:8080, paste the password, and use the xterm window to start amrvis2d or amrvis3d on your plt* directories.
Tip
On SLURM clusters with Apptainer, pull the image once with
apptainer pull amrvis-container.sif docker://ghcr.io/amrex-codes/amrvis-container:main, then use./launch_amrvis_browser_hpc.sh /path/to/plotfileson a compute node and follow the printed SSH tunnel instructions.
AMReX PlotfileTools
These are self-contained C++ programs (included with AMReX in the Tools/Plotfile subdirectory) that will output a 2D slice (axis-aligned), a 1D slice (axis-aligned), or compute a volume integral given an AMReX plotfile. This works as an alternative to yt and VisIt for basic tasks.
- To compute a volume integral, use fvolumesum.
- To compute a 2D slice plot (axis-aligned planes only), use fsnapshot.
- To compute a 1D slice (axis-aligned directions only, with output as ASCII), use fextract.
Other tools:
- fboxinfo prints out the indices of all the Boxes in a plotfile
- fcompare calculates the absolute and relative errors between plotfiles in L-inf norm
- fextrema calculates the minimum and maximum values of all variables in a plotfile
- fnan determines whether there are any NaNs in a plotfile
- ftime prints the simulation time of each plotfile
- fvarnames prints the names of all the variables in a given plotfile
yt
yt can load Quokka AMReX plotfiles through its Quokka frontend, which is now available in the main yt repository. Until a released yt version includes the frontend, install yt from the main branch with the optional Quokka dependencies:
pip install "yt[quokka] @ git+https://github.com/yt-project/yt.git"
After installation, load a plotfile with yt.load("plt00000") or point yt to any other Quokka plotfile directory. The upstream Quokka frontend documentation is available in the yt documentation source, and a tutorial notebook is available at README.ipynb.
Tip
One of the most useful things to do is to convert the data into a uniform-resolution NumPy array with the covering_grid function.
quick_plot script for batch processing
The quick_plot script in scripts/python/ wraps yt for batch visualization of Quokka outputs. It can process multiple snapshots and generate slice or projection plots. For usage details, see the documentation at the top of the script or run quick_plot -h.
yt-studio for web-based visualization
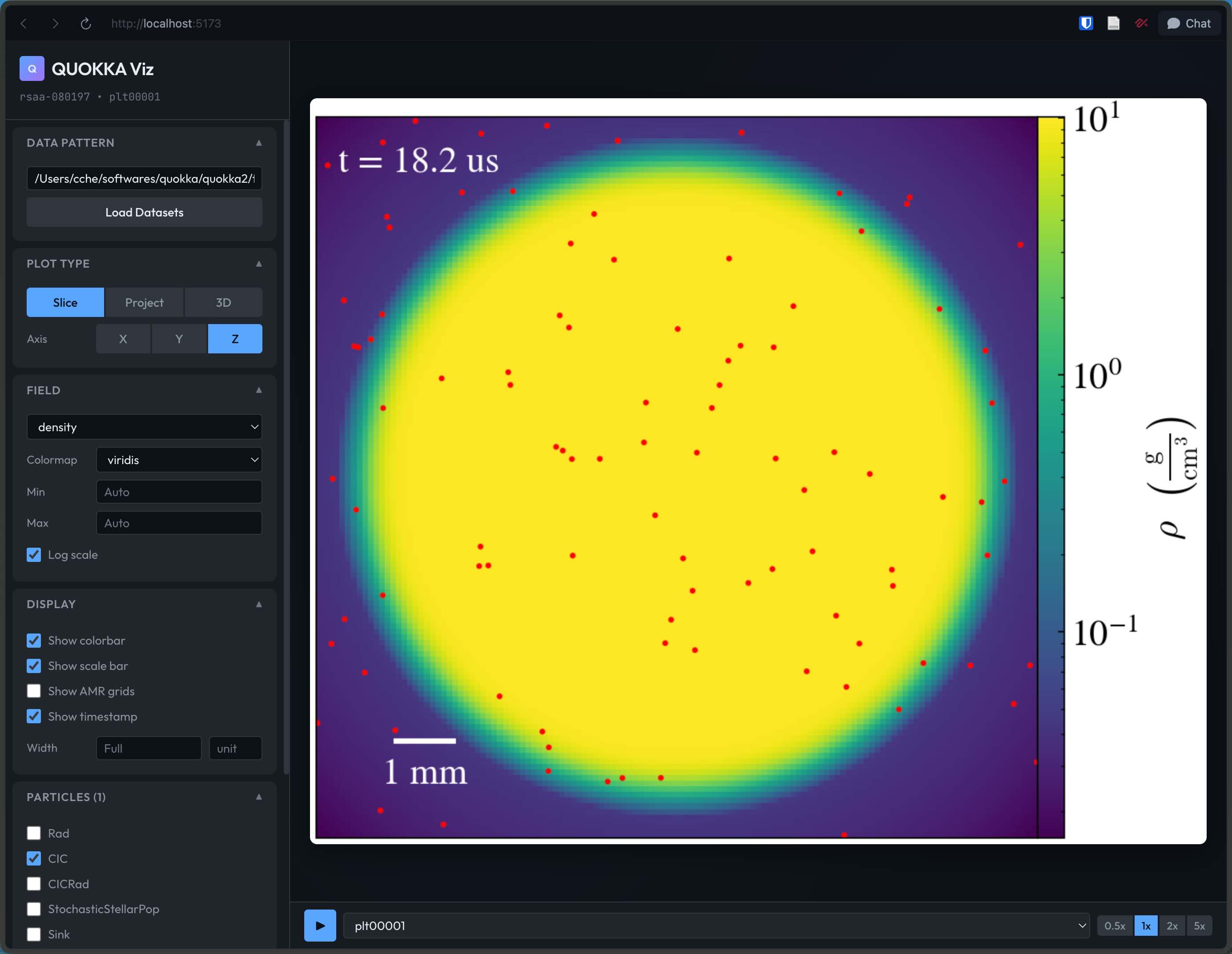
yt-studio provides a web interface and Python API for visualizing Quokka simulation data. Install it with pip install git+https://github.com/chongchonghe/yt-studio.git, then start the web interface with yt-studio and open http://localhost:5173 in your browser. It supports slice plots, projection plots, volume rendering, multiple colormaps, high-resolution export, optional particle overlays, and AMR grid annotations. For scripts and notebooks, use the QuokkaPlotter methods such as slice() and project().
VisIt
VisIt can read cell-centered output variables from AMReX plotfiles. Currently, there is no support for reading either face-centered variables or particles. (However, by default, cell-centered averages of face-centered variables are included in Quokka plotfiles.)
In order to read an individual plotfile, you can select the plt00000/Header file in VisIt’s Open dialog box.
If you want to read a timeseries of plotfiles, you can create a file with a .visit extension that lists the plt*/Header files, one per line, with the following command: :
ls -1 plt*/Header | tee plotfiles.visit
Then select plotfiles.visit in VisIt’s Open dialog box.
Warning
There are rendering bugs with unscaled box dimensions. Slices generally work. However, do not expect volume rendering to work when using, e.g. parsec-size boxes with cgs units.